You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001537_00657
You are here: Home > Sequence: MGYG000001537_00657
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Cellulomonas timonensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Actinomycetales; Cellulomonadaceae; Cellulomonas; Cellulomonas timonensis | |||||||||||
| CAZyme ID | MGYG000001537_00657 | |||||||||||
| CAZy Family | GH6 | |||||||||||
| CAZyme Description | Endoglucanase E-2 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 246021; End: 247595 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH6 | 57 | 380 | 3e-84 | 0.9523809523809523 |
| CBM2 | 422 | 510 | 1.6e-24 | 0.8712871287128713 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01341 | Glyco_hydro_6 | 1.03e-71 | 74 | 386 | 11 | 292 | Glycosyl hydrolases family 6. |
| COG5297 | CelA1 | 1.55e-35 | 88 | 501 | 165 | 540 | Cellulase/cellobiase CelA1 [Carbohydrate transport and metabolism]. |
| pfam00553 | CBM_2 | 1.55e-22 | 422 | 521 | 1 | 101 | Cellulose binding domain. Two tryptophan residues are involved in cellulose binding. Cellulose binding domain found in bacteria. |
| smart00637 | CBD_II | 2.71e-19 | 430 | 510 | 2 | 82 | CBD_II domain. |
| smart01063 | CBM49 | 8.41e-05 | 424 | 506 | 2 | 84 | Carbohydrate binding domain CBM49. This domain is found at the C terminal of cellulases and in vitro binding studies have shown it to binds to crystalline cellulose. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ALK04194.1 | 2.06e-238 | 48 | 523 | 55 | 534 |
| ARU50913.1 | 4.01e-238 | 33 | 523 | 40 | 533 |
| QNO37390.1 | 4.11e-231 | 8 | 522 | 7 | 514 |
| QEO14244.1 | 5.17e-228 | 26 | 522 | 22 | 519 |
| QIG39320.1 | 6.26e-174 | 22 | 522 | 15 | 525 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1UP3_A | 3.78e-45 | 48 | 409 | 10 | 288 | Structureof the endoglucanase Cel6 from Mycobacterium tuberculosis in complex with METHYL-CELLOBIOSYL-4-DEOXY-4-THIO-BETA-D-CELLOBIOSIDE at 1.6 angstrom [Mycobacterium tuberculosis H37Rv] |
| 1UOZ_A | 6.50e-45 | 48 | 409 | 31 | 309 | Structureof the endoglucanase Cel6 from Mycobacterium tuberculosis in complex with thiocellopentaose at 1.1 angstrom [Mycobacterium tuberculosis H37Rv] |
| 1UP0_A | 1.02e-44 | 48 | 409 | 10 | 288 | Structureof the endoglucanase Cel6 from Mycobacterium tuberculosis in complex with cellobiose at 1.75 angstrom [Mycobacterium tuberculosis H37Rv] |
| 1UP2_A | 1.97e-44 | 48 | 409 | 10 | 288 | Structureof the endoglucanase Cel6 from Mycobacterium tuberculosis in complex with glucose-isofagomine at 1.9 angstrom [Mycobacterium tuberculosis H37Rv] |
| 1TML_A | 8.34e-41 | 74 | 384 | 33 | 272 | CRYSTALSTRUCTURE OF THE CATALYTIC DOMAIN OF A THERMOPHILIC ENDOCELLULASE [Thermobifida fusca],2BOD_X Catalytic domain of endo-1,4-glucanase Cel6A from Thermobifida fusca in complex with methyl cellobiosyl-4-thio-beta-cellobioside [Thermobifida fusca] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P26414 | 1.93e-67 | 48 | 505 | 39 | 443 | Endoglucanase A OS=Thermobispora bispora OX=2006 GN=celA PE=3 SV=1 |
| P26222 | 6.08e-55 | 74 | 504 | 64 | 426 | Endoglucanase E-2 OS=Thermobifida fusca OX=2021 GN=celB PE=1 SV=2 |
| P07984 | 1.07e-34 | 41 | 409 | 168 | 443 | Endoglucanase A OS=Cellulomonas fimi OX=1708 GN=cenA PE=1 SV=1 |
| P13933 | 1.89e-33 | 74 | 400 | 107 | 359 | Endoglucanase 1 OS=Streptomyces sp. (strain KSM-9) OX=74575 GN=casA PE=1 SV=3 |
| P33682 | 8.11e-33 | 48 | 410 | 37 | 318 | Endoglucanase 1 OS=Streptomyces halstedii OX=1944 GN=celA1 PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
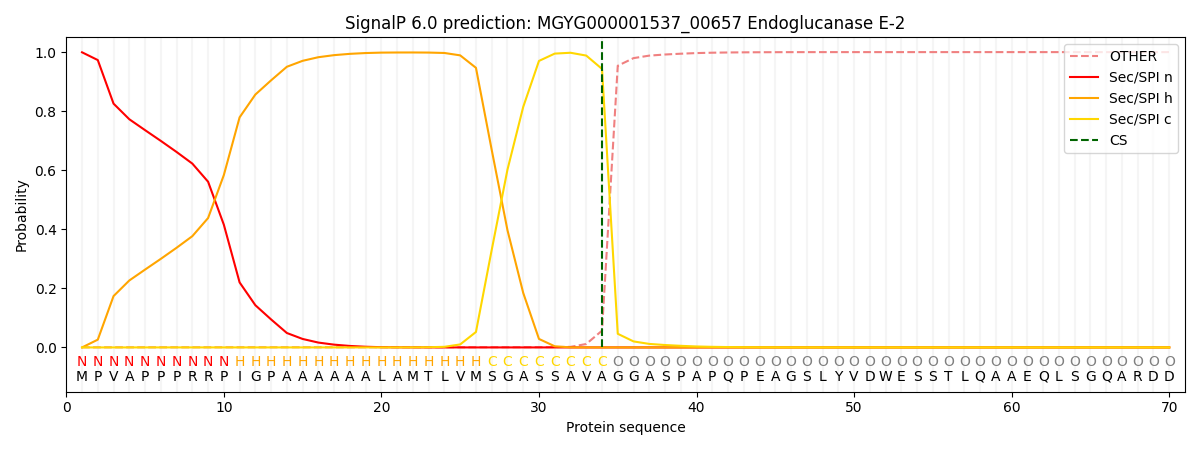
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001250 | 0.997412 | 0.000346 | 0.000545 | 0.000226 | 0.000184 |
