You are browsing environment: HUMAN GUT
MGYG000001539_01577
Basic Information
help
Species
Acutalibacter timonensis
Lineage
Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Acutalibacteraceae; Acutalibacter; Acutalibacter timonensis
CAZyme ID
MGYG000001539_01577
CAZy Family
GH55
CAZyme Description
hypothetical protein
CAZyme Property
Genome Property
Genome Assembly ID
Genome Size
Genome Type
Country
Continent
MGYG000001539
3671397
Isolate
not provided
not provided
Gene Location
Start: 683389;
End: 686238
Strand: -
No EC number prediction in MGYG000001539_01577.
Family
Start
End
Evalue
family coverage
GH55
116
574
4.3e-67
0.6432432432432432
Cdd ID
Domain
E-Value
qStart
qEnd
sStart
sEnd
Domain Description
pfam12708
Pectate_lyase_3
7.07e-17
348
480
1
144
Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28.
more
pfam12708
Pectate_lyase_3
4.00e-10
24
233
16
213
Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28.
more
COG5434
Pgu1
2.16e-04
349
399
83
137
Polygalacturonase [Carbohydrate transport and metabolism].
more
Hit ID
E-Value
Query Start
Query End
Hit Start
Hit End
Description
D4B0V1
7.61e-07
346
579
509
759
Probable glucan endo-1,3-beta-glucosidase ARB_02077 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_02077 PE=1 SV=1
more
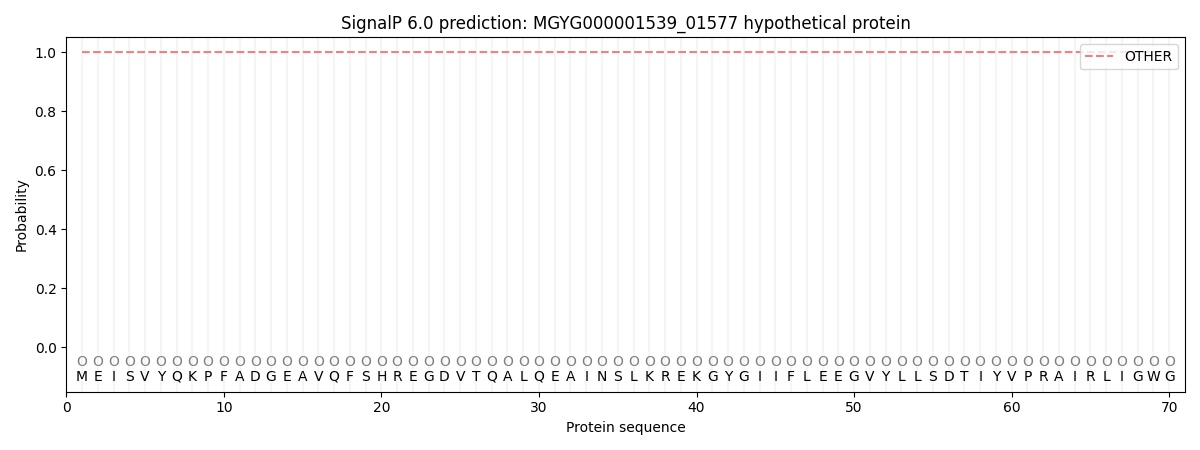
This protein is predicted as OTHER
Other
SP_Sec_SPI
LIPO_Sec_SPII
TAT_Tat_SPI
TATLIP_Sec_SPII
PILIN_Sec_SPIII
1.000072
0.000000
0.000000
0.000000
0.000000
0.000000
There is no transmembrane helices in MGYG000001539_01577.