You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001542_03746
You are here: Home > Sequence: MGYG000001542_03746
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
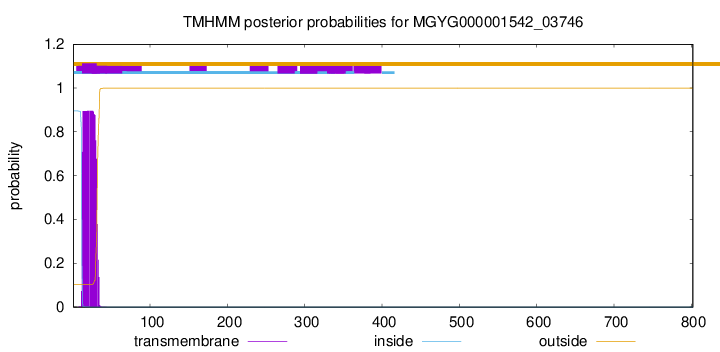
TMHMM annotations
Basic Information help
| Species | Paenibacillus_A sp900069005 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Paenibacillales; Paenibacillaceae; Paenibacillus_A; Paenibacillus_A sp900069005 | |||||||||||
| CAZyme ID | MGYG000001542_03746 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 467338; End: 469746 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 97 | 360 | 2.3e-119 | 0.9961977186311787 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00150 | Cellulase | 9.12e-26 | 118 | 359 | 23 | 268 | Cellulase (glycosyl hydrolase family 5). |
| COG2730 | BglC | 1.80e-19 | 64 | 370 | 14 | 372 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| cd00257 | Fascin | 8.03e-08 | 678 | 798 | 9 | 119 | Fascin-like domain; members include actin-bundling/crosslinking proteins facsin, histoactophilin and singed; identified in sea urchin, Drosophila, Xenopus, rodents, and humans; The fascin-like domain adopts a beta-trefoil topology and contains an internal threefold repeat; the fascin subgroup contains four copies of the domain; Structurally similar to fibroblast growth factor (FGF) |
| cd00257 | Fascin | 1.60e-07 | 440 | 557 | 10 | 117 | Fascin-like domain; members include actin-bundling/crosslinking proteins facsin, histoactophilin and singed; identified in sea urchin, Drosophila, Xenopus, rodents, and humans; The fascin-like domain adopts a beta-trefoil topology and contains an internal threefold repeat; the fascin subgroup contains four copies of the domain; Structurally similar to fibroblast growth factor (FGF) |
| COG3934 | COG3934 | 8.35e-06 | 123 | 337 | 31 | 253 | Endo-1,4-beta-mannosidase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AZS15177.1 | 0.0 | 1 | 565 | 1 | 565 |
| VEI36204.1 | 1.06e-226 | 15 | 562 | 14 | 550 |
| BCJ97694.1 | 3.00e-145 | 40 | 506 | 39 | 522 |
| QAA34430.1 | 4.30e-128 | 16 | 486 | 8 | 469 |
| AIC93076.1 | 1.90e-127 | 36 | 415 | 18 | 398 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1EQP_A | 7.96e-15 | 65 | 295 | 6 | 249 | Exo-b-(1,3)-glucanaseFrom Candida Albicans [Candida albicans] |
| 1CZ1_A | 1.42e-14 | 65 | 295 | 6 | 249 | Exo-b-(1,3)-glucanaseFrom Candida Albicans At 1.85 A Resolution [Candida albicans],1EQC_A Exo-b-(1,3)-glucanase From Candida Albicans In Complex With Castanospermine At 1.85 A [Candida albicans] |
| 4M80_A | 1.47e-14 | 65 | 295 | 11 | 254 | Thestructure of E292S glycosynthase variant of exo-1,3-beta-glucanase from Candida albicans at 1.85A resolution [Candida albicans SC5314],4M81_A The structure of E292S glycosynthase variant of exo-1,3-beta-glucanase from Candida albicans complexed with 1-fluoro-alpha-D-glucopyranoside (donor) and p-nitrophenyl beta-D-glucopyranoside (acceptor) at 1.86A resolution [Candida albicans SC5314],4M82_A The structure of E292S glycosynthase variant of exo-1,3-beta-glucanase from Candida albicans complexed with p-nitrophenyl-gentiobioside (product) at 1.6A resolution [Candida albicans SC5314] |
| 2PC8_A | 1.48e-14 | 65 | 295 | 12 | 255 | ChainA, Hypothetical protein XOG1 [Candida albicans] |
| 2PB1_A | 1.48e-14 | 65 | 295 | 12 | 255 | Exo-B-(1,3)-Glucanasefrom Candida Albicans in complex with unhydrolysed and covalently linked 2,4-dinitrophenyl-2-deoxy-2-fluoro-B-D-glucopyranoside at 1.9 A [Candida albicans] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| W8QRE4 | 4.22e-26 | 48 | 363 | 3 | 341 | Beta-xylosidase OS=Phanerodontia chrysosporium OX=2822231 GN=Xyl5 PE=1 SV=2 |
| B0XN12 | 6.66e-19 | 65 | 270 | 39 | 247 | Probable glucan 1,3-beta-glucosidase A OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=exgA PE=3 SV=1 |
| A1D4Q5 | 3.80e-18 | 65 | 270 | 39 | 247 | Probable glucan 1,3-beta-glucosidase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=exgA PE=3 SV=1 |
| Q4WK60 | 3.80e-18 | 65 | 261 | 39 | 238 | Probable glucan 1,3-beta-glucosidase A OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=exgA PE=3 SV=1 |
| B8N151 | 8.20e-18 | 68 | 332 | 31 | 300 | Probable glucan 1,3-beta-glucosidase A OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=exgA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
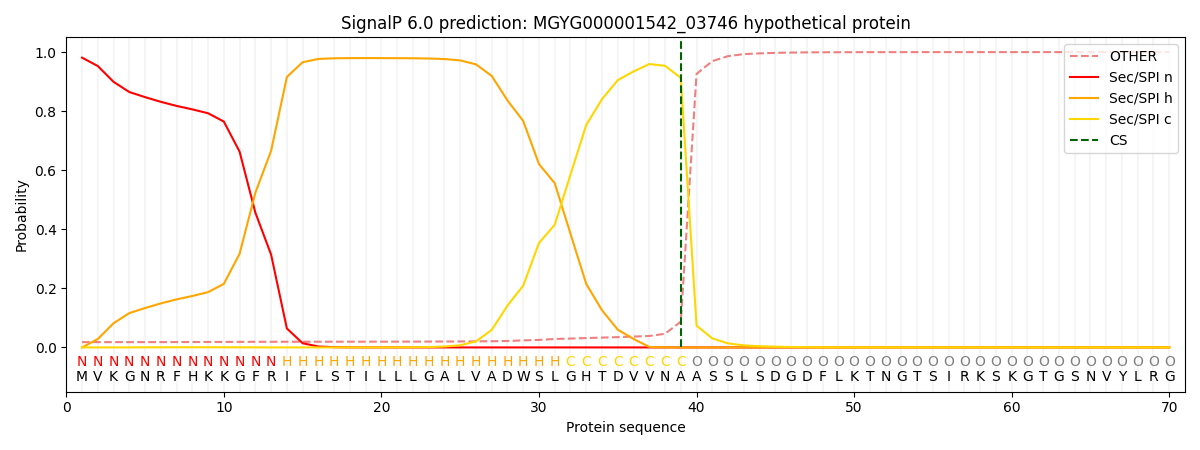
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.019480 | 0.978634 | 0.001041 | 0.000314 | 0.000240 | 0.000227 |