You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001548_00430
You are here: Home > Sequence: MGYG000001548_00430
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Paenibacillus_A tuaregi | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Paenibacillales; Paenibacillaceae; Paenibacillus_A; Paenibacillus_A tuaregi | |||||||||||
| CAZyme ID | MGYG000001548_00430 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 384096; End: 387392 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 429 | 768 | 2.6e-18 | 0.7963636363636364 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam02449 | Glyco_hydro_42 | 7.26e-08 | 447 | 698 | 10 | 260 | Beta-galactosidase. This group of beta-galactosidase enzymes belong to the glycosyl hydrolase 42 family. The enzyme catalyzes the hydrolysis of terminal, non-reducing terminal beta-D-galactosidase residues. |
| COG1874 | GanA | 1.96e-06 | 410 | 668 | 1 | 255 | Beta-galactosidase GanA [Carbohydrate transport and metabolism]. |
| cd03143 | A4_beta-galactosidase_middle_domain | 3.09e-04 | 817 | 932 | 1 | 119 | A4 beta-galactosidase middle domain: a type 1 glutamine amidotransferase (GATase1)-like domain. A4 beta-galactosidase middle domain: a type 1 glutamine amidotransferase (GATase1)-like domain. This group includes proteins similar to beta-galactosidase from Thermus thermophilus. Beta-Galactosidase hydrolyzes the beta-1,4-D-galactosidic linkage of lactose, as well as those of related chromogens, o-nitrophenyl-beta-D-galactopyranoside (ONP-Gal) and 5-bromo-4-chloro-3-indolyl-beta-D-galactoside (X-gal). This A4 beta-galactosidase middle domain lacks the catalytic triad of typical GATase1 domains. The reactive Cys residue found in the sharp turn between a beta strand and an alpha helix termed the nucleophile elbow in typical GATase1 domains is not conserved in this group. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QTH41876.1 | 0.0 | 6 | 1094 | 5 | 1029 |
| SDR98930.1 | 0.0 | 48 | 1094 | 34 | 1036 |
| QGG55714.1 | 0.0 | 43 | 1095 | 29 | 1036 |
| SQI53800.1 | 0.0 | 36 | 1095 | 24 | 1024 |
| QJC51567.1 | 0.0 | 43 | 1096 | 29 | 1037 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
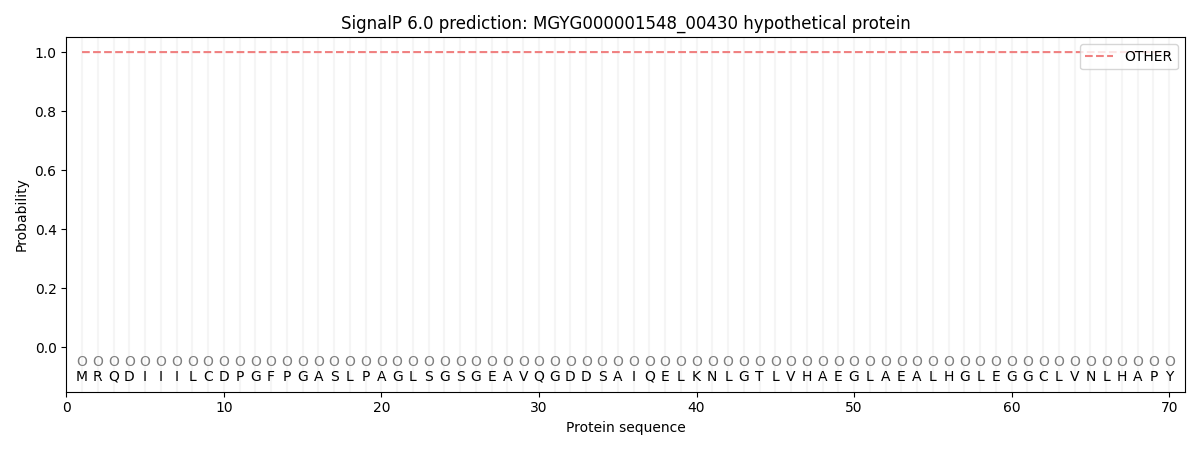
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000039 | 0.000001 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
