You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001548_01641
You are here: Home > Sequence: MGYG000001548_01641
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
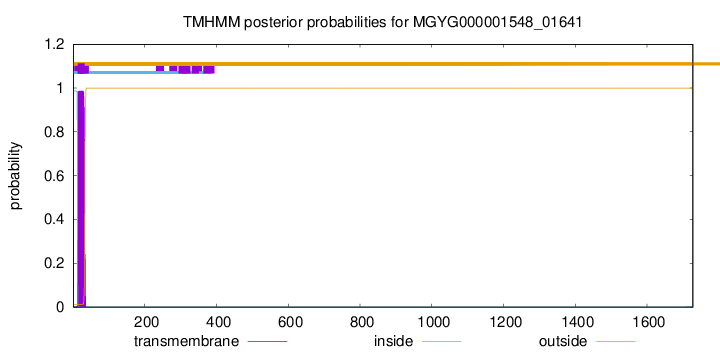
TMHMM annotations
Basic Information help
| Species | Paenibacillus_A tuaregi | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Paenibacillales; Paenibacillaceae; Paenibacillus_A; Paenibacillus_A tuaregi | |||||||||||
| CAZyme ID | MGYG000001548_01641 | |||||||||||
| CAZy Family | GH8 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1791941; End: 1797127 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01270 | Glyco_hydro_8 | 1.74e-22 | 1064 | 1394 | 31 | 317 | Glycosyl hydrolases family 8. |
| cd00839 | MPP_PAPs | 3.75e-19 | 475 | 722 | 22 | 254 | purple acid phosphatases of the metallophosphatase superfamily, metallophosphatase domain. Purple acid phosphatases (PAPs) belong to a diverse family of binuclear metallohydrolases that have been identified and characterized in plants, animals, and fungi. PAPs contain a binuclear metal center and their characteristic pink or purple color derives from a charge-transfer transition between a tyrosine residue and a chromophoric ferric ion within the binuclear center. PAPs catalyze the hydrolysis of a wide range of activated phosphoric acid mono- and di-esters and anhydrides. PAPs are distinguished from the other phosphatases by their insensitivity to L-(+) tartrate inhibition and are therefore also known as tartrate resistant acid phosphatases (TRAPs). While only a few copies of PAP-like genes are present in mammalian and fungal genomes, multiple copies are present in plant genomes. PAPs belong to the metallophosphatase (MPP) superfamily. MPPs are functionally diverse, but all share a conserved domain with an active site consisting of two metal ions (usually manganese, iron, or zinc) coordinated with octahedral geometry by a cage of histidine, aspartate, and asparagine residues. The MPP superfamily includes: Mre11/SbcD-like exonucleases, Dbr1-like RNA lariat debranching enzymes, YfcE-like phosphodiesterases, purple acid phosphatases (PAPs), YbbF-like UDP-2,3-diacylglucosamine hydrolases, and acid sphingomyelinases (ASMases). The conserved domain is a double beta-sheet sandwich with a di-metal active site made up of residues located at the C-terminal side of the sheets. This domain is thought to allow for productive metal coordination. |
| pfam16656 | Pur_ac_phosph_N | 7.74e-16 | 339 | 446 | 1 | 94 | Purple acid Phosphatase, N-terminal domain. This domain is found at the N-terminus of Purple acid phosphatase proteins. |
| NF033190 | inl_like_NEAT_1 | 6.01e-15 | 1525 | 1700 | 560 | 730 | NEAT domain-containing leucine-rich repeat protein. Members of this family have an N-terminal NEAT (near transporter) domain often associated with iron transport, followed by a leucine-rich repeat region with significant sequence similarity to the internalins of Listeria monocytogenes. However, since Bacillus cereus (from which this protein was described, in PMID:16978259) is not considered an intracellular pathogen, and the function may be iron transport rather than internalization, applying the name "internalin" to this family probably would be misleading. |
| cd00063 | FN3 | 1.64e-12 | 903 | 987 | 1 | 93 | Fibronectin type 3 domain; One of three types of internal repeats found in the plasma protein fibronectin. Its tenth fibronectin type III repeat contains an RGD cell recognition sequence in a flexible loop between 2 strands. Approximately 2% of all animal proteins contain the FN3 repeat; including extracellular and intracellular proteins, membrane spanning cytokine receptors, growth hormone receptors, tyrosine phosphatase receptors, and adhesion molecules. FN3-like domains are also found in bacterial glycosyl hydrolases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AIQ25865.1 | 0.0 | 52 | 1726 | 28 | 1696 |
| AZS18334.1 | 0.0 | 49 | 1721 | 58 | 1734 |
| AJS61487.1 | 0.0 | 40 | 1720 | 16 | 1704 |
| QNN60665.1 | 0.0 | 202 | 1493 | 10 | 1276 |
| QTH45936.1 | 8.54e-241 | 299 | 989 | 717 | 1383 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5XD0_A | 1.83e-135 | 996 | 1402 | 33 | 408 | ApoStructure of Beta-1,3-1,4-glucanase from Paenibacillus sp.X4 [Paenibacillus sp. X4],5XD0_B Apo Structure of Beta-1,3-1,4-glucanase from Paenibacillus sp.X4 [Paenibacillus sp. X4] |
| 7CJU_A | 1.08e-87 | 998 | 1409 | 11 | 396 | Crystalstructure of inactive form of chitosanase crystallized by ammonium sulfate [Bacillus sp. K17-2],7CJU_B Crystal structure of inactive form of chitosanase crystallized by ammonium sulfate [Bacillus sp. K17-2],7XGQ_A Chain A, chitosanase [Bacillus sp. K17-2],7XGQ_B Chain B, chitosanase [Bacillus sp. K17-2] |
| 1V5C_A | 5.05e-87 | 998 | 1403 | 5 | 384 | Thecrystal structure of the inactive form chitosanase from Bacillus sp. K17 at pH3.7 [Bacillus sp. (in: Bacteria)],1V5D_A The crystal structure of the active form chitosanase from Bacillus sp. K17 at pH6.4 [Bacillus sp. (in: Bacteria)],1V5D_B The crystal structure of the active form chitosanase from Bacillus sp. K17 at pH6.4 [Bacillus sp. (in: Bacteria)] |
| 1CEM_A | 1.57e-39 | 1017 | 1403 | 19 | 359 | ChainA, CELLULASE CELA (1,4-BETA-D-GLUCAN-GLUCANOHYDROLASE) [Acetivibrio thermocellus],1IS9_A Chain A, endoglucanase A [Acetivibrio thermocellus] |
| 1KWF_A | 3.28e-38 | 1017 | 1403 | 19 | 359 | ChainA, Endoglucanase A [Acetivibrio thermocellus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P19254 | 5.41e-136 | 996 | 1402 | 33 | 408 | Beta-glucanase OS=Niallia circulans OX=1397 GN=bgc PE=3 SV=1 |
| P29019 | 2.81e-85 | 980 | 1403 | 43 | 440 | Endoglucanase OS=Bacillus sp. (strain KSM-330) OX=72575 PE=1 SV=1 |
| A3DC29 | 4.89e-38 | 990 | 1414 | 25 | 402 | Endoglucanase A OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celA PE=1 SV=1 |
| C6CRV0 | 2.32e-37 | 1553 | 1722 | 1291 | 1460 | Endo-1,4-beta-xylanase A OS=Paenibacillus sp. (strain JDR-2) OX=324057 GN=xynA1 PE=1 SV=1 |
| P37699 | 2.98e-33 | 967 | 1400 | 2 | 389 | Endoglucanase C OS=Ruminiclostridium cellulolyticum (strain ATCC 35319 / DSM 5812 / JCM 6584 / H10) OX=394503 GN=celCCC PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
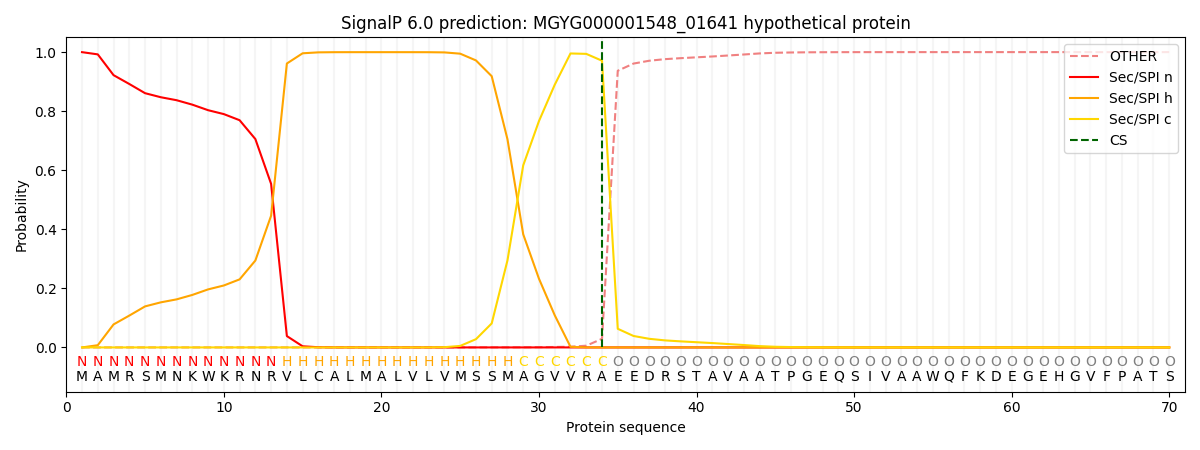
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000424 | 0.998504 | 0.000495 | 0.000210 | 0.000168 | 0.000148 |