You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001548_01951
You are here: Home > Sequence: MGYG000001548_01951
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Paenibacillus_A tuaregi | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Paenibacillales; Paenibacillaceae; Paenibacillus_A; Paenibacillus_A tuaregi | |||||||||||
| CAZyme ID | MGYG000001548_01951 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2079279; End: 2080721 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT2 | 62 | 331 | 4.5e-22 | 0.9956521739130435 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06423 | CESA_like | 3.40e-59 | 66 | 282 | 1 | 180 | CESA_like is the cellulose synthase superfamily. The cellulose synthase (CESA) superfamily includes a wide variety of glycosyltransferase family 2 enzymes that share the common characteristic of catalyzing the elongation of polysaccharide chains. The members include cellulose synthase catalytic subunit, chitin synthase, glucan biosynthesis protein and other families of CESA-like proteins. Cellulose synthase catalyzes the polymerization reaction of cellulose, an aggregate of unbranched polymers of beta-1,4-linked glucose residues in plants, most algae, some bacteria and fungi, and even some animals. In bacteria, algae and lower eukaryotes, there is a second unrelated type of cellulose synthase (Type II), which produces acylated cellulose, a derivative of cellulose. Chitin synthase catalyzes the incorporation of GlcNAc from substrate UDP-GlcNAc into chitin, which is a linear homopolymer of beta-(1,4)-linked GlcNAc residues and Glucan Biosynthesis protein catalyzes the elongation of beta-1,2 polyglucose chains of Glucan. |
| COG1215 | BcsA | 4.80e-43 | 15 | 434 | 12 | 384 | Glycosyltransferase, catalytic subunit of cellulose synthase and poly-beta-1,6-N-acetylglucosamine synthase [Cell motility]. |
| PRK11204 | PRK11204 | 4.45e-42 | 53 | 351 | 45 | 298 | N-glycosyltransferase; Provisional |
| PRK14583 | hmsR | 1.30e-25 | 62 | 358 | 75 | 326 | poly-beta-1,6 N-acetyl-D-glucosamine synthase. |
| pfam13641 | Glyco_tranf_2_3 | 1.27e-23 | 62 | 331 | 2 | 230 | Glycosyltransferase like family 2. Members of this family of prokaryotic proteins include putative glucosyltransferase, which are involved in bacterial capsule biosynthesis. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AWB45147.1 | 1.49e-302 | 20 | 480 | 2 | 462 |
| AZK49032.1 | 6.97e-287 | 20 | 475 | 2 | 457 |
| QCT02414.1 | 4.09e-283 | 11 | 473 | 2 | 464 |
| ANY76433.1 | 1.69e-282 | 20 | 475 | 2 | 457 |
| ACX64312.1 | 1.09e-281 | 18 | 475 | 3 | 460 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5HEA_A | 1.10e-06 | 64 | 209 | 7 | 120 | CgTstructure in hexamer [Streptococcus parasanguinis FW213],5HEA_B CgT structure in hexamer [Streptococcus parasanguinis FW213],5HEA_C CgT structure in hexamer [Streptococcus parasanguinis FW213],5HEC_A CgT structure in dimer [Streptococcus parasanguinis FW213],5HEC_B CgT structure in dimer [Streptococcus parasanguinis FW213] |
| 2Z86_A | 4.82e-06 | 49 | 184 | 360 | 472 | Crystalstructure of chondroitin polymerase from Escherichia coli strain K4 (K4CP) complexed with UDP-GlcUA and UDP [Escherichia coli],2Z86_B Crystal structure of chondroitin polymerase from Escherichia coli strain K4 (K4CP) complexed with UDP-GlcUA and UDP [Escherichia coli],2Z86_C Crystal structure of chondroitin polymerase from Escherichia coli strain K4 (K4CP) complexed with UDP-GlcUA and UDP [Escherichia coli],2Z86_D Crystal structure of chondroitin polymerase from Escherichia coli strain K4 (K4CP) complexed with UDP-GlcUA and UDP [Escherichia coli] |
| 2Z87_A | 4.82e-06 | 49 | 184 | 359 | 471 | Crystalstructure of chondroitin polymerase from Escherichia coli strain K4 (K4CP) complexed with UDP-GalNAc and UDP [Escherichia coli],2Z87_B Crystal structure of chondroitin polymerase from Escherichia coli strain K4 (K4CP) complexed with UDP-GalNAc and UDP [Escherichia coli] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8GLC5 | 4.43e-26 | 56 | 347 | 41 | 286 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus epidermidis OX=1282 GN=icaA PE=3 SV=1 |
| Q5HKQ0 | 1.52e-25 | 56 | 347 | 41 | 286 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus epidermidis (strain ATCC 35984 / RP62A) OX=176279 GN=icaA PE=1 SV=1 |
| Q7A351 | 2.71e-23 | 46 | 347 | 32 | 286 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus aureus (strain N315) OX=158879 GN=icaA PE=3 SV=1 |
| Q6GDD8 | 2.71e-23 | 46 | 347 | 32 | 286 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus aureus (strain MRSA252) OX=282458 GN=icaA PE=3 SV=1 |
| Q5HCN1 | 2.71e-23 | 46 | 347 | 32 | 286 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus aureus (strain COL) OX=93062 GN=icaA PE=3 SV=1 |
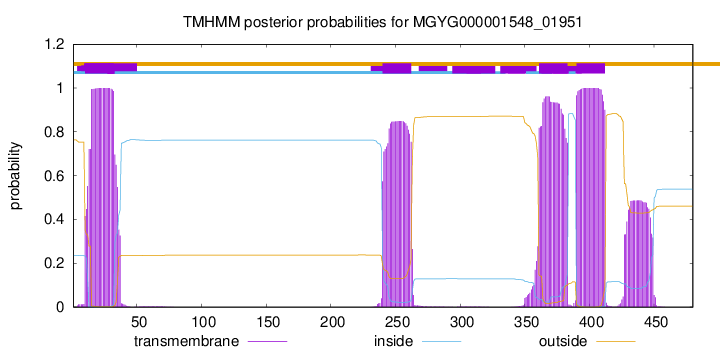
SignalP and Lipop Annotations help
This protein is predicted as OTHER
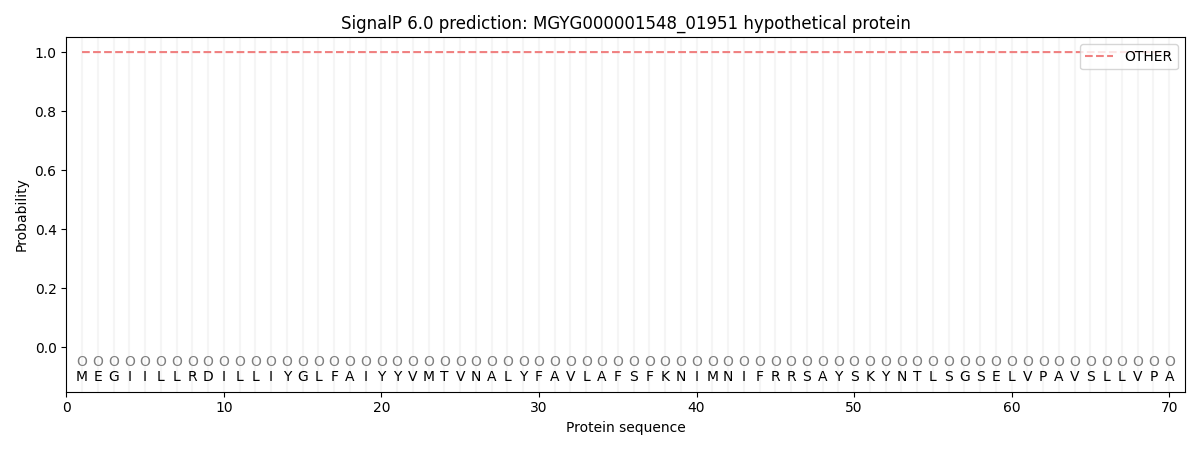
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000024 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |