You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001554_00240
You are here: Home > Sequence: MGYG000001554_00240
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Corynebacterium phoceense | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Mycobacteriales; Mycobacteriaceae; Corynebacterium; Corynebacterium phoceense | |||||||||||
| CAZyme ID | MGYG000001554_00240 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | Beta-hexosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 247700; End: 248884 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 137 | 352 | 2.5e-34 | 0.9351851851851852 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1472 | BglX | 2.99e-49 | 91 | 384 | 18 | 305 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| pfam00933 | Glyco_hydro_3 | 4.86e-48 | 82 | 386 | 7 | 312 | Glycosyl hydrolase family 3 N terminal domain. |
| PRK05337 | PRK05337 | 1.84e-36 | 126 | 352 | 48 | 278 | beta-hexosaminidase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QPK80426.1 | 5.79e-152 | 69 | 384 | 82 | 397 |
| APT93375.1 | 5.69e-147 | 66 | 386 | 70 | 388 |
| QQU82600.1 | 1.21e-130 | 73 | 393 | 57 | 381 |
| AMO92002.1 | 1.29e-130 | 74 | 393 | 67 | 387 |
| QRQ67786.1 | 1.71e-130 | 73 | 389 | 57 | 374 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5IOB_A | 7.94e-100 | 74 | 379 | 25 | 335 | Crystalstructure of beta-N-acetylglucosaminidase-like protein from Corynebacterium glutamicum [Corynebacterium glutamicum ATCC 13032],5IOB_B Crystal structure of beta-N-acetylglucosaminidase-like protein from Corynebacterium glutamicum [Corynebacterium glutamicum ATCC 13032],5IOB_C Crystal structure of beta-N-acetylglucosaminidase-like protein from Corynebacterium glutamicum [Corynebacterium glutamicum ATCC 13032],5IOB_D Crystal structure of beta-N-acetylglucosaminidase-like protein from Corynebacterium glutamicum [Corynebacterium glutamicum ATCC 13032],5IOB_E Crystal structure of beta-N-acetylglucosaminidase-like protein from Corynebacterium glutamicum [Corynebacterium glutamicum ATCC 13032],5IOB_F Crystal structure of beta-N-acetylglucosaminidase-like protein from Corynebacterium glutamicum [Corynebacterium glutamicum ATCC 13032],5IOB_G Crystal structure of beta-N-acetylglucosaminidase-like protein from Corynebacterium glutamicum [Corynebacterium glutamicum ATCC 13032],5IOB_H Crystal structure of beta-N-acetylglucosaminidase-like protein from Corynebacterium glutamicum [Corynebacterium glutamicum ATCC 13032] |
| 4YYF_A | 2.61e-62 | 82 | 373 | 46 | 341 | ChainA, Beta-N-acetylhexosaminidase [Mycolicibacterium smegmatis MC2 155],4YYF_B Chain B, Beta-N-acetylhexosaminidase [Mycolicibacterium smegmatis MC2 155],4YYF_C Chain C, Beta-N-acetylhexosaminidase [Mycolicibacterium smegmatis MC2 155] |
| 6GFV_A | 2.20e-57 | 82 | 376 | 23 | 322 | Mtuberculosis LpqI [Mycobacterium tuberculosis H37Rv],6GFV_B M tuberculosis LpqI [Mycobacterium tuberculosis H37Rv] |
| 5G1M_A | 1.10e-23 | 122 | 352 | 64 | 293 | Crystalstructure of NagZ from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5G1M_B Crystal structure of NagZ from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5G2M_A Crystal structure of NagZ from Pseudomonas aeruginosa in complex with N-acetylglucosamine [Pseudomonas aeruginosa PAO1],5G2M_B Crystal structure of NagZ from Pseudomonas aeruginosa in complex with N-acetylglucosamine [Pseudomonas aeruginosa PAO1],5G3R_A Crystal Structure Of Nagz From Pseudomonas Aeruginosa In Complex With N-acetylglucosamine And L-ala-1,6-anhydromurnac [Pseudomonas aeruginosa PAO1],5G3R_B Crystal Structure Of Nagz From Pseudomonas Aeruginosa In Complex With N-acetylglucosamine And L-ala-1,6-anhydromurnac [Pseudomonas aeruginosa PAO1],5G5K_A Crystal structure of NagZ from Pseudomonas aeruginosa in complex with the inhibitor 2-acetamido-1,2-dideoxynojirimycin [Pseudomonas aeruginosa PAO1],5G5K_B Crystal structure of NagZ from Pseudomonas aeruginosa in complex with the inhibitor 2-acetamido-1,2-dideoxynojirimycin [Pseudomonas aeruginosa PAO1] |
| 6JTI_A | 2.25e-22 | 99 | 359 | 65 | 330 | Crystalstructure of native NagZ from Neisseria gonorrhoeae [Neisseria gonorrhoeae],6JTI_B Crystal structure of native NagZ from Neisseria gonorrhoeae [Neisseria gonorrhoeae],6JTI_C Crystal structure of native NagZ from Neisseria gonorrhoeae [Neisseria gonorrhoeae],6JTI_D Crystal structure of native NagZ from Neisseria gonorrhoeae [Neisseria gonorrhoeae],6JTI_E Crystal structure of native NagZ from Neisseria gonorrhoeae [Neisseria gonorrhoeae],6JTI_F Crystal structure of native NagZ from Neisseria gonorrhoeae [Neisseria gonorrhoeae],6JTJ_A Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-acetylglucosamine [Neisseria gonorrhoeae],6JTJ_B Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-acetylglucosamine [Neisseria gonorrhoeae],6JTJ_C Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-acetylglucosamine [Neisseria gonorrhoeae],6JTJ_D Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-acetylglucosamine [Neisseria gonorrhoeae],6JTJ_E Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-acetylglucosamine [Neisseria gonorrhoeae],6JTJ_F Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-acetylglucosamine [Neisseria gonorrhoeae],6JTK_A Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-trifluoroacetyl-D-glucosamine [Neisseria gonorrhoeae],6JTK_B Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-trifluoroacetyl-D-glucosamine [Neisseria gonorrhoeae],6JTK_C Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-trifluoroacetyl-D-glucosamine [Neisseria gonorrhoeae],6JTK_D Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-trifluoroacetyl-D-glucosamine [Neisseria gonorrhoeae],6JTK_E Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-trifluoroacetyl-D-glucosamine [Neisseria gonorrhoeae],6JTK_F Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-trifluoroacetyl-D-glucosamine [Neisseria gonorrhoeae],6JTL_A Crystal structure of NagZ from Neisseria gonorrhoeae in complex with zinc ion [Neisseria gonorrhoeae],6JTL_B Crystal structure of NagZ from Neisseria gonorrhoeae in complex with zinc ion [Neisseria gonorrhoeae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| L7N6B0 | 3.87e-56 | 82 | 376 | 66 | 365 | Beta-hexosaminidase LpqI OS=Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) OX=83332 GN=lpqI PE=1 SV=1 |
| A0A0H3M1P5 | 1.07e-55 | 82 | 376 | 66 | 365 | Beta-hexosaminidase LpqI OS=Mycobacterium bovis (strain BCG / Pasteur 1173P2) OX=410289 GN=lpqI PE=3 SV=1 |
| B0U3L0 | 2.44e-27 | 126 | 368 | 46 | 292 | Beta-hexosaminidase OS=Xylella fastidiosa (strain M12) OX=405440 GN=nagZ PE=3 SV=1 |
| Q9PAZ0 | 4.64e-27 | 126 | 368 | 46 | 292 | Beta-hexosaminidase OS=Xylella fastidiosa (strain 9a5c) OX=160492 GN=nagZ PE=3 SV=1 |
| B2FPW9 | 1.22e-26 | 126 | 352 | 46 | 276 | Beta-hexosaminidase OS=Stenotrophomonas maltophilia (strain K279a) OX=522373 GN=nagZ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
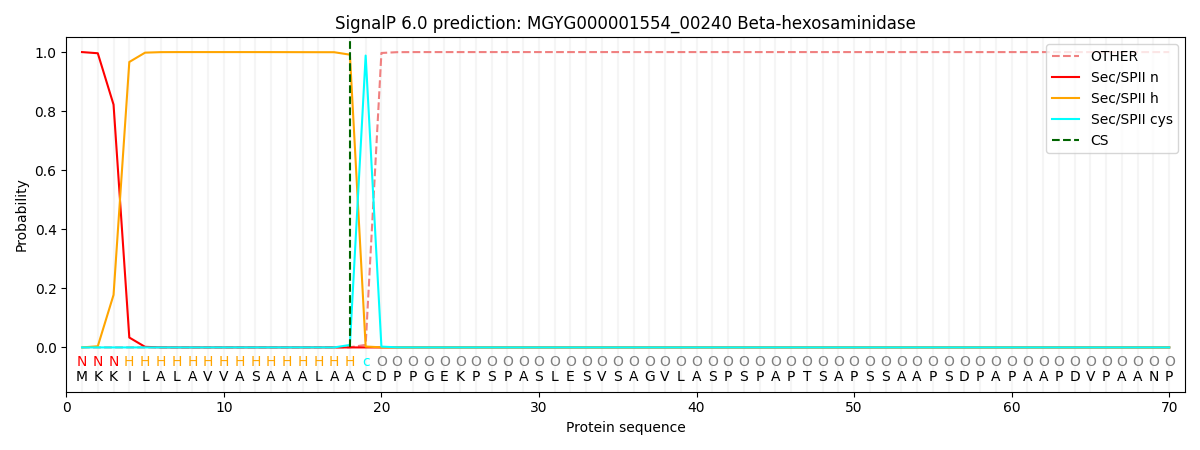
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000001 | 1.000071 | 0.000000 | 0.000000 | 0.000000 |
