You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001554_00533
You are here: Home > Sequence: MGYG000001554_00533
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Corynebacterium phoceense | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Mycobacteriales; Mycobacteriaceae; Corynebacterium; Corynebacterium phoceense | |||||||||||
| CAZyme ID | MGYG000001554_00533 | |||||||||||
| CAZy Family | CE1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 547100; End: 548734 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE1 | 123 | 445 | 1.5e-38 | 0.986784140969163 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG0627 | FrmB | 1.49e-35 | 121 | 451 | 45 | 316 | S-formylglutathione hydrolase FrmB [Defense mechanisms]. |
| pfam00756 | Esterase | 1.09e-16 | 123 | 436 | 1 | 239 | Putative esterase. This family contains Esterase D. However it is not clear if all members of the family have the same function. This family is related to the pfam00135 family. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QVI99079.1 | 3.49e-81 | 115 | 449 | 64 | 355 |
| AOX06778.1 | 3.86e-80 | 104 | 448 | 60 | 377 |
| AKK10240.1 | 1.45e-73 | 129 | 445 | 75 | 367 |
| QXW04485.1 | 1.84e-60 | 114 | 448 | 59 | 347 |
| BCN74918.1 | 2.96e-60 | 76 | 448 | 24 | 340 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4H18_A | 1.97e-93 | 104 | 451 | 59 | 365 | Threedimensional structure of corynomycoloyl tranferase C [Corynebacterium glutamicum ATCC 13032],4H18_B Three dimensional structure of corynomycoloyl tranferase C [Corynebacterium glutamicum ATCC 13032],4H18_C Three dimensional structure of corynomycoloyl tranferase C [Corynebacterium glutamicum ATCC 13032],4H18_D Three dimensional structure of corynomycoloyl tranferase C [Corynebacterium glutamicum ATCC 13032] |
| 6SX4_AAA | 1.42e-18 | 118 | 448 | 64 | 344 | ChainAAA, Protein PS1 [Corynebacterium glutamicum],6SX4_BBB Chain BBB, Protein PS1 [Corynebacterium glutamicum],6SX4_CCC Chain CCC, Protein PS1 [Corynebacterium glutamicum],6SX4_DDD Chain DDD, Protein PS1 [Corynebacterium glutamicum] |
| 1SFR_A | 3.05e-16 | 116 | 453 | 9 | 289 | ChainA, Antigen 85-A [Mycobacterium tuberculosis],1SFR_B Chain B, Antigen 85-A [Mycobacterium tuberculosis],1SFR_C Chain C, Antigen 85-A [Mycobacterium tuberculosis] |
| 7MYG_A | 1.89e-14 | 116 | 436 | 4 | 265 | ChainA, Diacylglycerol acyltransferase [Mycobacterium tuberculosis],7MYG_B Chain B, Diacylglycerol acyltransferase [Mycobacterium tuberculosis] |
| 1DQZ_A | 1.94e-14 | 116 | 436 | 6 | 267 | ChainA, PROTEIN (ANTIGEN 85-C) [Mycobacterium tuberculosis],1DQZ_B Chain B, PROTEIN (ANTIGEN 85-C) [Mycobacterium tuberculosis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P0C1D7 | 2.17e-20 | 118 | 454 | 107 | 391 | Protein PS1 OS=Corynebacterium melassecola OX=41643 GN=csp1 PE=4 SV=1 |
| P0C1D6 | 2.88e-20 | 118 | 454 | 107 | 391 | Protein PS1 OS=Corynebacterium glutamicum (strain ATCC 13032 / DSM 20300 / BCRC 11384 / JCM 1318 / LMG 3730 / NCIMB 10025) OX=196627 GN=csp1 PE=1 SV=1 |
| O06052 | 5.73e-16 | 144 | 432 | 74 | 309 | Diacylglycerol acyltransferase/mycolyltransferase Ag85A OS=Mycobacterium gordonae OX=1778 GN=fbpA PE=3 SV=1 |
| P9WQP3 | 2.48e-15 | 116 | 453 | 51 | 331 | Diacylglycerol acyltransferase/mycolyltransferase Ag85A OS=Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) OX=83332 GN=fbpA PE=1 SV=1 |
| P9WQP2 | 2.48e-15 | 116 | 453 | 51 | 331 | Diacylglycerol acyltransferase/mycolyltransferase Ag85A OS=Mycobacterium tuberculosis (strain CDC 1551 / Oshkosh) OX=83331 GN=fbpA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
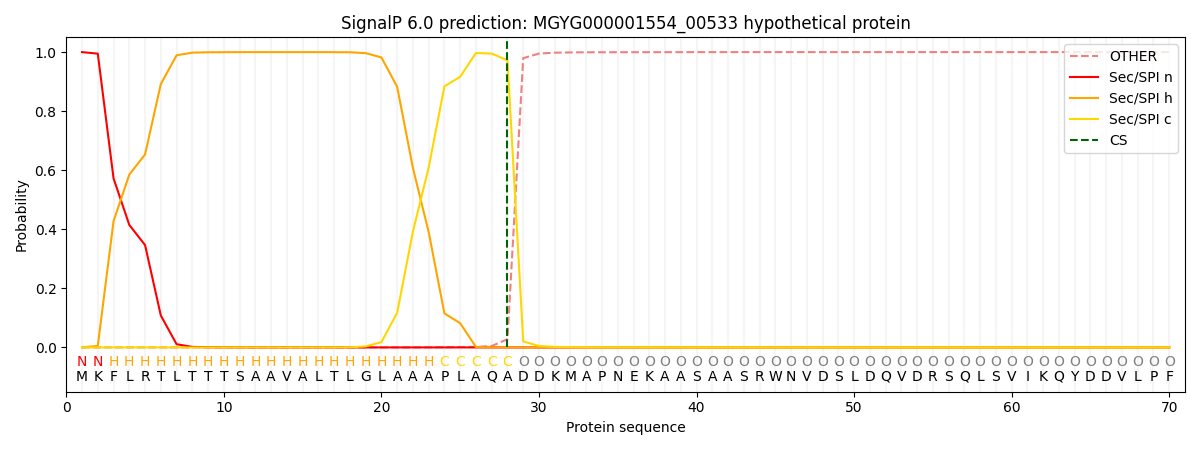
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000302 | 0.998924 | 0.000200 | 0.000233 | 0.000168 | 0.000145 |
