You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001561_00923
You are here: Home > Sequence: MGYG000001561_00923
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Caecibacter massiliensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_C; Negativicutes; Veillonellales; Megasphaeraceae; Caecibacter; Caecibacter massiliensis | |||||||||||
| CAZyme ID | MGYG000001561_00923 | |||||||||||
| CAZy Family | GT83 | |||||||||||
| CAZyme Description | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 966833; End: 968458 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT83 | 4 | 464 | 6.4e-86 | 0.8851851851851852 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1807 | ArnT | 1.45e-38 | 4 | 432 | 7 | 442 | 4-amino-4-deoxy-L-arabinose transferase or related glycosyltransferase of PMT family [Cell wall/membrane/envelope biogenesis]. |
| PRK13279 | arnT | 2.24e-20 | 1 | 311 | 2 | 317 | lipid IV(A) 4-amino-4-deoxy-L-arabinosyltransferase. |
| pfam13231 | PMT_2 | 2.18e-10 | 60 | 219 | 1 | 157 | Dolichyl-phosphate-mannose-protein mannosyltransferase. This family contains members that are not captured by pfam02366. |
| COG4745 | COG4745 | 2.55e-04 | 46 | 338 | 46 | 383 | Predicted membrane-bound mannosyltransferase [General function prediction only]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AXB83023.1 | 0.0 | 1 | 541 | 1 | 541 |
| AXL21032.1 | 2.88e-231 | 1 | 536 | 1 | 536 |
| ALG41393.1 | 7.56e-197 | 1 | 534 | 1 | 532 |
| CCC72525.1 | 1.07e-196 | 1 | 534 | 1 | 532 |
| AVO73980.1 | 1.07e-196 | 1 | 534 | 1 | 532 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5EZM_A | 9.02e-28 | 27 | 311 | 54 | 348 | CrystalStructure of ArnT from Cupriavidus metallidurans in the apo state [Cupriavidus metallidurans CH34],5F15_A Crystal Structure of ArnT from Cupriavidus metallidurans bound to Undecaprenyl phosphate [Cupriavidus metallidurans CH34] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B7LM74 | 1.94e-25 | 2 | 334 | 3 | 339 | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase OS=Escherichia fergusonii (strain ATCC 35469 / DSM 13698 / CCUG 18766 / IAM 14443 / JCM 21226 / LMG 7866 / NBRC 102419 / NCTC 12128 / CDC 0568-73) OX=585054 GN=arnT PE=3 SV=2 |
| O67601 | 2.21e-24 | 7 | 335 | 5 | 337 | Uncharacterized protein aq_1704 OS=Aquifex aeolicus (strain VF5) OX=224324 GN=aq_1704 PE=3 SV=1 |
| Q6D2F3 | 6.67e-24 | 29 | 338 | 32 | 346 | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase OS=Pectobacterium atrosepticum (strain SCRI 1043 / ATCC BAA-672) OX=218491 GN=arnT PE=3 SV=1 |
| C6DAW3 | 8.92e-24 | 29 | 338 | 32 | 346 | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase OS=Pectobacterium carotovorum subsp. carotovorum (strain PC1) OX=561230 GN=arnT PE=3 SV=1 |
| B2VBI7 | 1.18e-23 | 29 | 336 | 29 | 342 | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase OS=Erwinia tasmaniensis (strain DSM 17950 / CFBP 7177 / CIP 109463 / NCPPB 4357 / Et1/99) OX=465817 GN=arnT PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
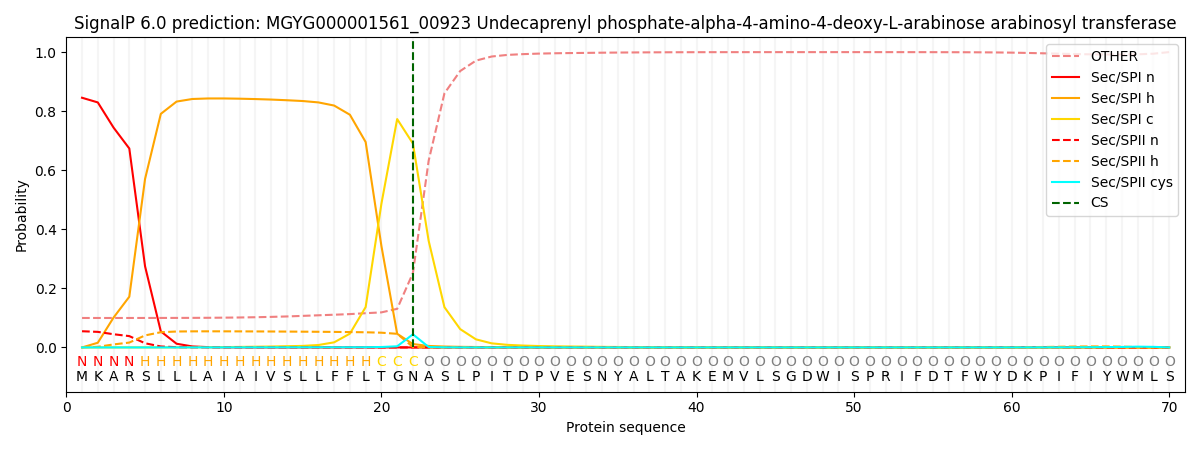
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.112894 | 0.827432 | 0.057289 | 0.000708 | 0.000658 | 0.000968 |
TMHMM Annotations download full data without filtering help
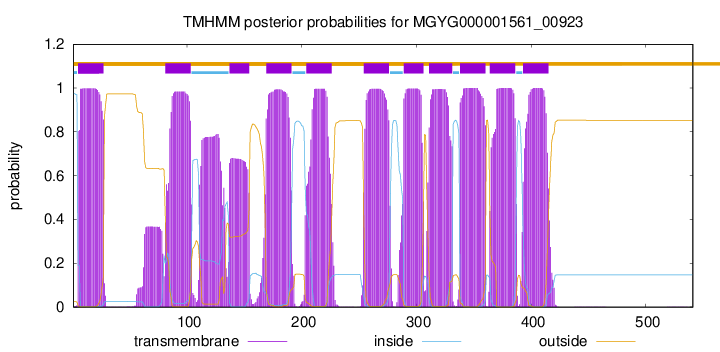

| start | end |
|---|---|
| 5 | 27 |
| 81 | 103 |
| 137 | 154 |
| 169 | 191 |
| 204 | 226 |
| 254 | 276 |
| 289 | 306 |
| 311 | 331 |
| 338 | 360 |
| 364 | 386 |
| 393 | 415 |
