You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001614_01404
You are here: Home > Sequence: MGYG000001614_01404
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | CAG-521 sp002329575 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Burkholderiales; Burkholderiaceae; CAG-521; CAG-521 sp002329575 | |||||||||||
| CAZyme ID | MGYG000001614_01404 | |||||||||||
| CAZy Family | GH8 | |||||||||||
| CAZyme Description | Endoglucanase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 5780; End: 6859 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK11097 | PRK11097 | 4.63e-133 | 21 | 351 | 24 | 364 | cellulase. |
| COG3405 | BcsZ | 1.61e-70 | 22 | 345 | 26 | 350 | Endo-1,4-beta-D-glucanase Y [Carbohydrate transport and metabolism]. |
| pfam01270 | Glyco_hydro_8 | 4.87e-48 | 21 | 341 | 3 | 321 | Glycosyl hydrolases family 8. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QDA53874.1 | 4.17e-147 | 2 | 351 | 1 | 358 |
| QQS88762.1 | 1.26e-141 | 19 | 355 | 25 | 363 |
| ANU67169.1 | 4.58e-124 | 1 | 355 | 1 | 352 |
| QQQ96024.1 | 4.58e-124 | 1 | 355 | 1 | 352 |
| QDA53782.1 | 4.06e-100 | 22 | 355 | 71 | 411 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4Q2B_A | 4.17e-96 | 21 | 351 | 3 | 339 | Thecrystal structure of an endo-1,4-D-glucanase from Pseudomonas putida KT2440 [Pseudomonas putida KT2440],4Q2B_B The crystal structure of an endo-1,4-D-glucanase from Pseudomonas putida KT2440 [Pseudomonas putida KT2440],4Q2B_C The crystal structure of an endo-1,4-D-glucanase from Pseudomonas putida KT2440 [Pseudomonas putida KT2440],4Q2B_D The crystal structure of an endo-1,4-D-glucanase from Pseudomonas putida KT2440 [Pseudomonas putida KT2440],4Q2B_E The crystal structure of an endo-1,4-D-glucanase from Pseudomonas putida KT2440 [Pseudomonas putida KT2440],4Q2B_F The crystal structure of an endo-1,4-D-glucanase from Pseudomonas putida KT2440 [Pseudomonas putida KT2440] |
| 5CD2_A | 2.32e-87 | 22 | 349 | 26 | 361 | Thecrystal structure of endo-1,4-D-glucanase from Vibrio fischeri ES114 [Aliivibrio fischeri ES114] |
| 7F81_A | 1.76e-84 | 19 | 351 | 5 | 339 | ChainA, Glucanase [Enterobacter sp. CJF-002],7F81_B Chain B, Glucanase [Enterobacter sp. CJF-002],7F81_C Chain C, Glucanase [Enterobacter sp. CJF-002],7F81_D Chain D, Glucanase [Enterobacter sp. CJF-002] |
| 3QXQ_A | 3.98e-84 | 19 | 351 | 1 | 335 | Structureof the bacterial cellulose synthase subunit Z in complex with cellopentaose [Escherichia coli K-12],3QXQ_B Structure of the bacterial cellulose synthase subunit Z in complex with cellopentaose [Escherichia coli K-12],3QXQ_C Structure of the bacterial cellulose synthase subunit Z in complex with cellopentaose [Escherichia coli K-12],3QXQ_D Structure of the bacterial cellulose synthase subunit Z in complex with cellopentaose [Escherichia coli K-12] |
| 7F82_A | 4.98e-84 | 19 | 351 | 5 | 339 | ChainA, Glucanase [Enterobacter sp. CJF-002],7F82_B Chain B, Glucanase [Enterobacter sp. CJF-002],7F82_C Chain C, Glucanase [Enterobacter sp. CJF-002],7F82_D Chain D, Glucanase [Enterobacter sp. CJF-002] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8ZLB7 | 2.32e-86 | 10 | 351 | 14 | 357 | Endoglucanase OS=Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) OX=99287 GN=bcsZ PE=3 SV=1 |
| Q8X5L9 | 6.36e-86 | 6 | 351 | 8 | 356 | Endoglucanase OS=Escherichia coli O157:H7 OX=83334 GN=bcsZ PE=3 SV=1 |
| Q8Z289 | 1.85e-85 | 10 | 351 | 14 | 357 | Endoglucanase OS=Salmonella typhi OX=90370 GN=bcsZ PE=3 SV=1 |
| P37651 | 1.43e-84 | 9 | 351 | 12 | 356 | Endoglucanase OS=Escherichia coli (strain K12) OX=83333 GN=bcsZ PE=1 SV=1 |
| P58935 | 4.97e-71 | 22 | 351 | 33 | 371 | Endoglucanase OS=Xanthomonas axonopodis pv. citri (strain 306) OX=190486 GN=bcsZ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
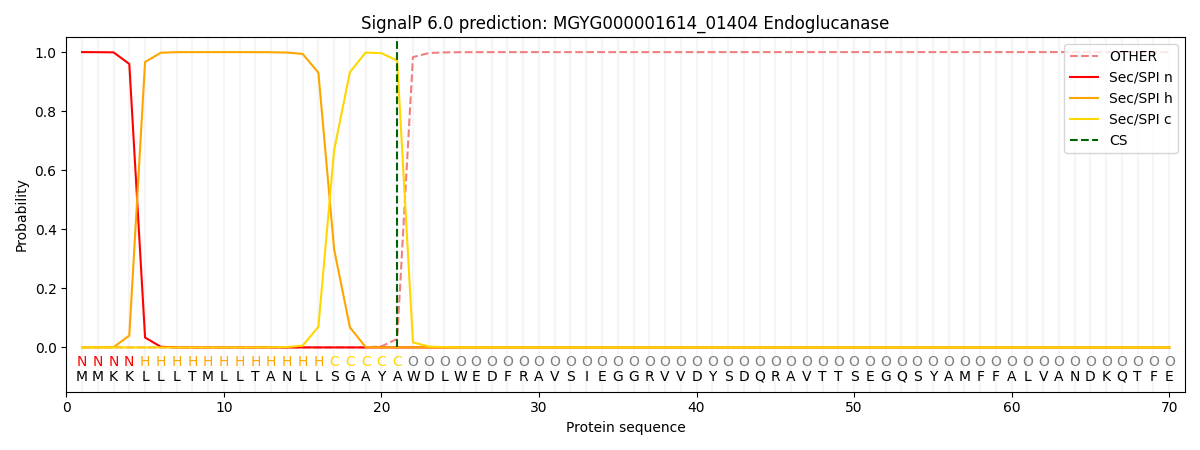
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000290 | 0.999050 | 0.000179 | 0.000159 | 0.000150 | 0.000141 |
