You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001615_02972
You are here: Home > Sequence: MGYG000001615_02972
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Eisenbergiella sp900548905 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Eisenbergiella; Eisenbergiella sp900548905 | |||||||||||
| CAZyme ID | MGYG000001615_02972 | |||||||||||
| CAZy Family | GH140 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 97112; End: 98443 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH140 | 13 | 437 | 6.2e-134 | 0.9902912621359223 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam13204 | DUF4038 | 1.23e-124 | 20 | 345 | 1 | 318 | Protein of unknown function (DUF4038). A family of putative cellulases. |
| pfam12904 | Collagen_bind_2 | 3.21e-14 | 354 | 442 | 2 | 91 | Putative collagen-binding domain of a collagenase. This domain is likely to be the collagen-binding domain of a family of bacterial collagenase enzymes. It is the C-terminal part of the Structure 3kzs (information derived from TOPSAN). |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QOV18626.1 | 1.07e-151 | 11 | 442 | 3 | 434 |
| AUO19127.1 | 5.69e-143 | 30 | 442 | 22 | 432 |
| ANW98439.1 | 3.58e-137 | 15 | 442 | 4 | 438 |
| ANX00976.1 | 3.58e-137 | 15 | 442 | 4 | 438 |
| AGC68049.1 | 3.58e-137 | 15 | 442 | 4 | 438 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5MSY_A | 9.09e-119 | 9 | 442 | 12 | 444 | Glycosidehydrolase BT_1012 [Bacteroides thetaiotaomicron VPI-5482],5MSY_B Glycoside hydrolase BT_1012 [Bacteroides thetaiotaomicron VPI-5482],5MSY_C Glycoside hydrolase BT_1012 [Bacteroides thetaiotaomicron VPI-5482] |
| 3KZS_A | 4.76e-113 | 9 | 442 | 12 | 444 | Crystalstructure of glycosyl hydrolase family 5 (NP_809925.1) from BACTEROIDES THETAIOTAOMICRON VPI-5482 at 2.10 A resolution [Bacteroides thetaiotaomicron VPI-5482],3KZS_B Crystal structure of glycosyl hydrolase family 5 (NP_809925.1) from BACTEROIDES THETAIOTAOMICRON VPI-5482 at 2.10 A resolution [Bacteroides thetaiotaomicron VPI-5482],3KZS_C Crystal structure of glycosyl hydrolase family 5 (NP_809925.1) from BACTEROIDES THETAIOTAOMICRON VPI-5482 at 2.10 A resolution [Bacteroides thetaiotaomicron VPI-5482],3KZS_D Crystal structure of glycosyl hydrolase family 5 (NP_809925.1) from BACTEROIDES THETAIOTAOMICRON VPI-5482 at 2.10 A resolution [Bacteroides thetaiotaomicron VPI-5482] |
| 4QFU_A | 2.06e-105 | 9 | 442 | 25 | 460 | ChainA, glycoside hydrolase family 5 [Phocaeicola vulgatus ATCC 8482],4QFU_B Chain B, glycoside hydrolase family 5 [Phocaeicola vulgatus ATCC 8482],4QFU_C Chain C, glycoside hydrolase family 5 [Phocaeicola vulgatus ATCC 8482],4QFU_D Chain D, glycoside hydrolase family 5 [Phocaeicola vulgatus ATCC 8482],4QFU_E Chain E, glycoside hydrolase family 5 [Phocaeicola vulgatus ATCC 8482],4QFU_F Chain F, glycoside hydrolase family 5 [Phocaeicola vulgatus ATCC 8482],4QFU_G Chain G, glycoside hydrolase family 5 [Phocaeicola vulgatus ATCC 8482],4QFU_H Chain H, glycoside hydrolase family 5 [Phocaeicola vulgatus ATCC 8482],4QFU_I Chain I, glycoside hydrolase family 5 [Phocaeicola vulgatus ATCC 8482],4QFU_J Chain J, glycoside hydrolase family 5 [Phocaeicola vulgatus ATCC 8482],4QFU_K Chain K, glycoside hydrolase family 5 [Phocaeicola vulgatus ATCC 8482],4QFU_L Chain L, glycoside hydrolase family 5 [Phocaeicola vulgatus ATCC 8482] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
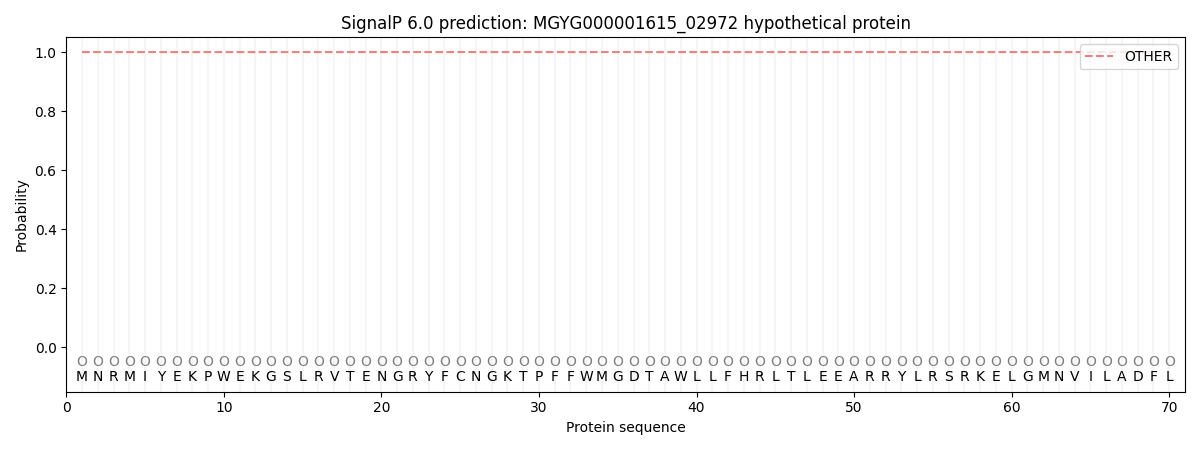
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000055 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
