You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001639_03264
You are here: Home > Sequence: MGYG000001639_03264
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Haloferax massiliensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Archaea; Halobacteriota; Halobacteria; Halobacteriales; Haloferacaceae; Haloferax; Haloferax massiliensis | |||||||||||
| CAZyme ID | MGYG000001639_03264 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 202821; End: 204587 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH16 | 66 | 291 | 3.1e-31 | 0.9841269841269841 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd08023 | GH16_laminarinase_like | 1.19e-29 | 65 | 291 | 1 | 235 | Laminarinase, member of the glycosyl hydrolase family 16. Laminarinase, also known as glucan endo-1,3-beta-D-glucosidase, is a glycosyl hydrolase family 16 member that hydrolyzes 1,3-beta-D-glucosidic linkages in 1,3-beta-D-glucans such as laminarins, curdlans, paramylons, and pachymans, with very limited action on mixed-link (1,3-1,4-)-beta-D-glucans. |
| cd00413 | Glyco_hydrolase_16 | 4.58e-26 | 67 | 291 | 1 | 210 | glycosyl hydrolase family 16. The O-Glycosyl hydrolases are a widespread group of enzymes that hydrolyse the glycosidic bond between two or more carbohydrates, or between a carbohydrate and a non-carbohydrate moiety. A glycosyl hydrolase classification system based on sequence similarity has led to the definition of more than 95 different families inlcuding glycosyl hydrolase family 16. Family 16 includes lichenase, xyloglucan endotransglycosylase (XET), beta-agarase, kappa-carrageenase, endo-beta-1,3-glucanase, endo-beta-1,3-1,4-glucanase, and endo-beta-galactosidase, all of which have a conserved jelly roll fold with a deep active site channel harboring the catalytic residues. |
| cd02178 | GH16_beta_agarase | 2.63e-24 | 56 | 291 | 14 | 257 | Beta-agarase, member of glycosyl hydrolase family 16. Beta-agarase is a glycosyl hydrolase family 16 (GH16) member that hydrolyzes the internal beta-1,4-linkage of agarose, a hydrophilic polysaccharide found in the cell wall of Rhodophyceaea, marine red algae. Agarose is a linear chain of galactose units linked by alternating L-alpha-1,3- and D-beta-1,4-linkages that are additionally modified by a 3,6-anhydro-bridge. Agarose forms thermo-reversible gels that are widely used in the food industry or as a laboratory medium. While beta-agarases are also found in two other families derived from the sequence-based classification of glycosyl hydrolases (GH50, and GH86) the GH16 members are most abundant. This domain adopts a curved beta-sandwich conformation, with a tunnel-shaped active site cavity, referred to as a jellyroll fold. |
| COG2273 | BglS | 2.64e-17 | 47 | 289 | 27 | 261 | Beta-glucanase, GH16 family [Carbohydrate transport and metabolism]. |
| pfam00722 | Glyco_hydro_16 | 3.94e-14 | 100 | 250 | 5 | 134 | Glycosyl hydrolases family 16. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QCJ47429.1 | 3.34e-139 | 2 | 587 | 3 | 598 |
| AGB39278.1 | 8.51e-30 | 46 | 297 | 41 | 307 |
| ATW88142.1 | 1.58e-26 | 61 | 290 | 15 | 261 |
| QNU67712.1 | 2.67e-21 | 55 | 297 | 356 | 596 |
| QSG01745.1 | 1.32e-20 | 29 | 294 | 18 | 286 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2VY0_A | 4.27e-17 | 60 | 291 | 12 | 259 | TheX-ray structure of endo-beta-1,3-glucanase from Pyrococcus furiosus [Pyrococcus furiosus],2VY0_B The X-ray structure of endo-beta-1,3-glucanase from Pyrococcus furiosus [Pyrococcus furiosus] |
| 4DFS_A | 2.61e-09 | 59 | 292 | 16 | 263 | Structureof the catalytic domain of an endo-1,3-beta-glucanase (laminarinase) from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],4DFS_B Structure of the catalytic domain of an endo-1,3-beta-glucanase (laminarinase) from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1] |
| 3AZX_A | 4.94e-09 | 59 | 291 | 8 | 254 | Crystalstructure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3AZX_B Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3AZY_A Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3AZY_B Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3AZY_C Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3AZY_D Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3AZZ_A Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 in complex with gluconolactone [Thermotoga maritima MSB8],3AZZ_B Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 in complex with gluconolactone [Thermotoga maritima MSB8],3AZZ_C Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 in complex with gluconolactone [Thermotoga maritima MSB8],3AZZ_D Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 in complex with gluconolactone [Thermotoga maritima MSB8],3B00_A Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 in complex with cetyltrimethylammonium bromide [Thermotoga maritima MSB8],3B00_B Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 in complex with cetyltrimethylammonium bromide [Thermotoga maritima MSB8],3B00_C Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 in complex with cetyltrimethylammonium bromide [Thermotoga maritima MSB8],3B00_D Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 in complex with cetyltrimethylammonium bromide [Thermotoga maritima MSB8],3B01_A Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3B01_B Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3B01_C Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3B01_D Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8] |
| 3I4I_A | 5.88e-09 | 72 | 246 | 29 | 190 | Crystalstructure of a prokaryotic beta-1,3-1,4-glucanase (lichenase) derived from a mouse hindgut metagenome [uncultured murine large bowel bacterium BAC 14],3I4I_B Crystal structure of a prokaryotic beta-1,3-1,4-glucanase (lichenase) derived from a mouse hindgut metagenome [uncultured murine large bowel bacterium BAC 14] |
| 6T2R_AAA | 1.10e-08 | 60 | 289 | 28 | 259 | ChainAAA, Beta-glycosidase [Bacteroides caccae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P23903 | 3.08e-12 | 63 | 291 | 425 | 679 | Glucan endo-1,3-beta-glucosidase A1 OS=Niallia circulans OX=1397 GN=glcA PE=1 SV=1 |
| O33680 | 5.38e-09 | 137 | 291 | 320 | 457 | Endo-1,3-1,4-beta-glycanase ExsH OS=Rhizobium meliloti (strain 1021) OX=266834 GN=exsH PE=1 SV=1 |
| P37073 | 1.32e-08 | 72 | 289 | 46 | 248 | Beta-glucanase OS=Brevibacillus brevis OX=1393 GN=bglBB PE=1 SV=1 |
| P33693 | 1.52e-07 | 64 | 293 | 37 | 233 | Endo-1,3-1,4-beta-glycanase ExoK OS=Rhizobium meliloti (strain 1021) OX=266834 GN=exoK PE=1 SV=1 |
| Q9Z3Q2 | 2.04e-07 | 75 | 291 | 257 | 457 | Endo-1,3-1,4-beta-glycanase EglC OS=Rhizobium meliloti (strain 1021) OX=266834 GN=eglC PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as TAT
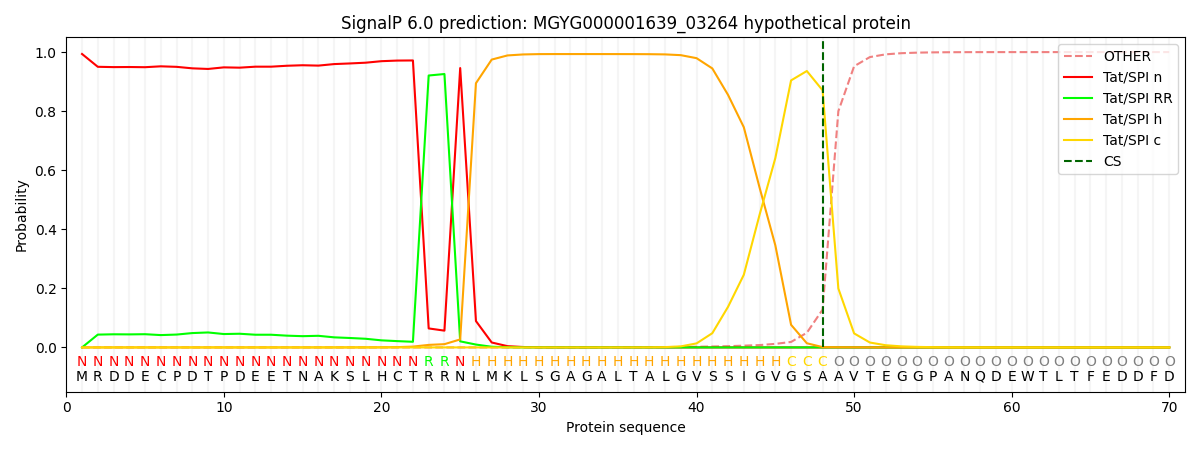
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000003 | 0.000003 | 0.000001 | 0.993620 | 0.006357 | 0.000000 |
