You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001643_00817
You are here: Home > Sequence: MGYG000001643_00817
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Muribaculaceae; CAG-485; | |||||||||||
| CAZyme ID | MGYG000001643_00817 | |||||||||||
| CAZy Family | GH115 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 18778; End: 21129 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH115 | 22 | 777 | 3.7e-226 | 0.9956958393113343 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam15979 | Glyco_hydro_115 | 1.17e-180 | 159 | 503 | 1 | 334 | Glycosyl hydrolase family 115. Glyco_hydro_115 is a family of glycoside hydrolases likely to have the activity of xylan a-1,2-glucuronidase, EC:3.2.1.131, or a-(4-O-methyl)-glucuronidase EC:3.2.1.-. |
| cd05833 | Ribosomal_P2 | 1.61e-04 | 54 | 94 | 8 | 48 | Ribosomal protein P2. This subfamily represents the eukaryotic large ribosomal protein P2. Eukaryotic P1 and P2 are functionally equivalent to the bacterial protein L7/L12, but are not homologous to L7/L12. P2 is located in the L12 stalk, with proteins P1, P0, L11, and 28S rRNA. P1 and P2 are the only proteins in the ribosome to occur as multimers, always appearing as sets of heterodimers. Recent data indicate that eukaryotes have four copies (two heterodimers), while most archaeal species contain six copies of L12p (three homodimers). Bacteria may have four or six copies of L7/L12 (two or three homodimers) depending on the species. Experiments using S. cerevisiae P1 and P2 indicate that P1 proteins are positioned more internally with limited reactivity in the C-terminal domains, while P2 proteins seem to be more externally located and are more likely to interact with other cellular components. In lower eukaryotes, P1 and P2 are further subdivided into P1A, P1B, P2A, and P2B, which form P1A/P2B and P1B/P2A heterodimers. Some plants have a third P-protein, called P3, which is not homologous to P1 and P2. In humans, P1 and P2 are strongly autoimmunogenic. They play a significant role in the etiology and pathogenesis of systemic lupus erythema (SLE). In addition, the ribosome-inactivating protein trichosanthin (TCS) interacts with human P0, P1, and P2, with its primary binding site in the C-terminal region of P2. TCS inactivates the ribosome by depurinating a specific adenine in the sarcin-ricin loop of 28S rRNA. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| EDY97203.1 | 0.0 | 21 | 779 | 20 | 782 |
| ALJ39737.1 | 0.0 | 22 | 780 | 25 | 780 |
| QDO67466.1 | 0.0 | 12 | 780 | 10 | 778 |
| BCA52290.1 | 0.0 | 22 | 780 | 25 | 780 |
| QUT73086.1 | 0.0 | 22 | 780 | 25 | 780 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5BY3_A | 0.0 | 22 | 780 | 19 | 774 | Anovel family GH115 4-O-Methyl-alpha-glucuronidase, BtGH115A, with specificity for decorated arabinogalactans [Bacteroides thetaiotaomicron VPI-5482] |
| 4C90_A | 8.89e-91 | 18 | 622 | 39 | 640 | Evidencethat GH115 alpha-glucuronidase activity is dependent on conformational flexibility [Bacteroides ovatus],4C90_B Evidence that GH115 alpha-glucuronidase activity is dependent on conformational flexibility [Bacteroides ovatus],4C91_A Evidence that GH115 alpha-glucuronidase activity is dependent on conformational flexibility [Bacteroides ovatus],4C91_B Evidence that GH115 alpha-glucuronidase activity is dependent on conformational flexibility [Bacteroides ovatus] |
| 4ZMH_A | 3.68e-88 | 71 | 618 | 68 | 606 | Crystalstructure of a five-domain GH115 alpha-Glucuronidase from the Marine Bacterium Saccharophagus degradans 2-40T [Saccharophagus degradans 2-40],4ZMH_B Crystal structure of a five-domain GH115 alpha-Glucuronidase from the Marine Bacterium Saccharophagus degradans 2-40T [Saccharophagus degradans 2-40] |
| 7PUG_A | 1.28e-87 | 25 | 625 | 20 | 628 | ChainA, xylan alpha-1,2-glucuronidase [uncultured bacterium] |
| 7PXQ_A | 2.43e-87 | 25 | 625 | 19 | 627 | ChainA, xylan alpha-1,2-glucuronidase [uncultured bacterium] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
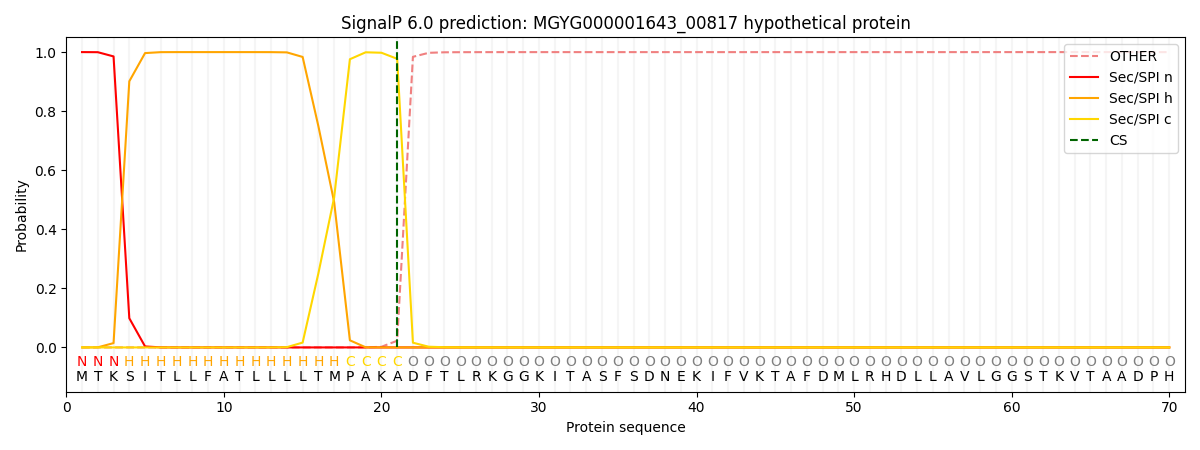
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000234 | 0.999071 | 0.000204 | 0.000158 | 0.000158 | 0.000152 |
