You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001652_01131
You are here: Home > Sequence: MGYG000001652_01131
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | V9D3004 sp002349525 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; V9D3004; V9D3004 sp002349525 | |||||||||||
| CAZyme ID | MGYG000001652_01131 | |||||||||||
| CAZy Family | GH10 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 89; End: 3010 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 610 | 971 | 1.9e-76 | 0.9504950495049505 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00633 | Glyco_10 | 8.13e-70 | 681 | 969 | 14 | 263 | Glycosyl hydrolase family 10. |
| pfam00331 | Glyco_hydro_10 | 7.76e-64 | 599 | 971 | 1 | 310 | Glycosyl hydrolase family 10. |
| COG3693 | XynA | 2.93e-54 | 589 | 967 | 17 | 335 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
| PRK00708 | PRK00708 | 5.33e-10 | 148 | 245 | 104 | 202 | twin-arginine translocase subunit TatB. |
| NF033913 | fibronec_FbpA | 6.23e-10 | 118 | 245 | 160 | 301 | LPXTG-anchored fibronectin-binding protein FbpA. FbpA, a fibronectin-binding protein described in Streptococcus pyogenes, has a YSIRK-type (crosswall-targeting) signal peptide and a C-terminal LPXTG motif for covalent attachment to the cell wall. It is unrelated to the PavA-like protein from Streptococcus gordonii (see BlastRule NBR009716) that was given the identical name, so the phase LPXTG-anchored is added to the protein name for clarity. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AFU34339.1 | 5.11e-109 | 521 | 973 | 88 | 520 |
| ABX41884.1 | 1.91e-103 | 562 | 973 | 134 | 515 |
| BCJ93966.1 | 3.48e-95 | 573 | 973 | 141 | 513 |
| CDM69886.1 | 2.99e-91 | 598 | 972 | 203 | 557 |
| CBK75021.1 | 7.93e-91 | 579 | 973 | 12 | 382 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5OFJ_A | 1.75e-42 | 626 | 972 | 37 | 338 | Crystalstructure of N-terminal domain of bifunctional CbXyn10C [Caldicellulosiruptor bescii DSM 6725] |
| 6D5C_A | 5.70e-42 | 609 | 972 | 32 | 350 | Structureof Caldicellulosiruptor danielii GH10 module of glycoside hydrolase WP_045175321 [Caldicellulosiruptor danielii],6D5C_B Structure of Caldicellulosiruptor danielii GH10 module of glycoside hydrolase WP_045175321 [Caldicellulosiruptor danielii],6D5C_C Structure of Caldicellulosiruptor danielii GH10 module of glycoside hydrolase WP_045175321 [Caldicellulosiruptor danielii] |
| 7NL2_A | 8.11e-42 | 595 | 972 | 8 | 340 | ChainA, Beta-xylanase [Pseudothermotoga thermarum DSM 5069],7NL2_B Chain B, Beta-xylanase [Pseudothermotoga thermarum DSM 5069] |
| 5OFK_A | 1.09e-41 | 626 | 972 | 37 | 338 | Crystalstructure of CbXyn10C variant E140Q/E248Q complexed with xyloheptaose [Caldicellulosiruptor bescii DSM 6725],5OFL_A Crystal structure of CbXyn10C variant E140Q/E248Q complexed with cellohexaose [Caldicellulosiruptor bescii DSM 6725] |
| 2W5F_A | 4.53e-38 | 610 | 966 | 194 | 521 | ChainA, ENDO-1,4-BETA-XYLANASE Y [Acetivibrio thermocellus],2W5F_B Chain B, ENDO-1,4-BETA-XYLANASE Y [Acetivibrio thermocellus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P23551 | 1.10e-75 | 607 | 973 | 46 | 383 | Endo-1,4-beta-xylanase A OS=Butyrivibrio fibrisolvens OX=831 GN=xynA PE=3 SV=1 |
| P29126 | 3.80e-40 | 627 | 966 | 655 | 946 | Bifunctional endo-1,4-beta-xylanase XylA OS=Ruminococcus flavefaciens OX=1265 GN=xynA PE=3 SV=1 |
| Q60042 | 1.91e-39 | 595 | 970 | 362 | 686 | Endo-1,4-beta-xylanase A OS=Thermotoga neapolitana OX=2337 GN=xynA PE=1 SV=1 |
| Q60037 | 1.92e-39 | 588 | 970 | 359 | 690 | Endo-1,4-beta-xylanase A OS=Thermotoga maritima (strain ATCC 43589 / DSM 3109 / JCM 10099 / NBRC 100826 / MSB8) OX=243274 GN=xynA PE=1 SV=1 |
| P10474 | 3.97e-38 | 626 | 972 | 71 | 372 | Endoglucanase/exoglucanase B OS=Caldicellulosiruptor saccharolyticus OX=44001 GN=celB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
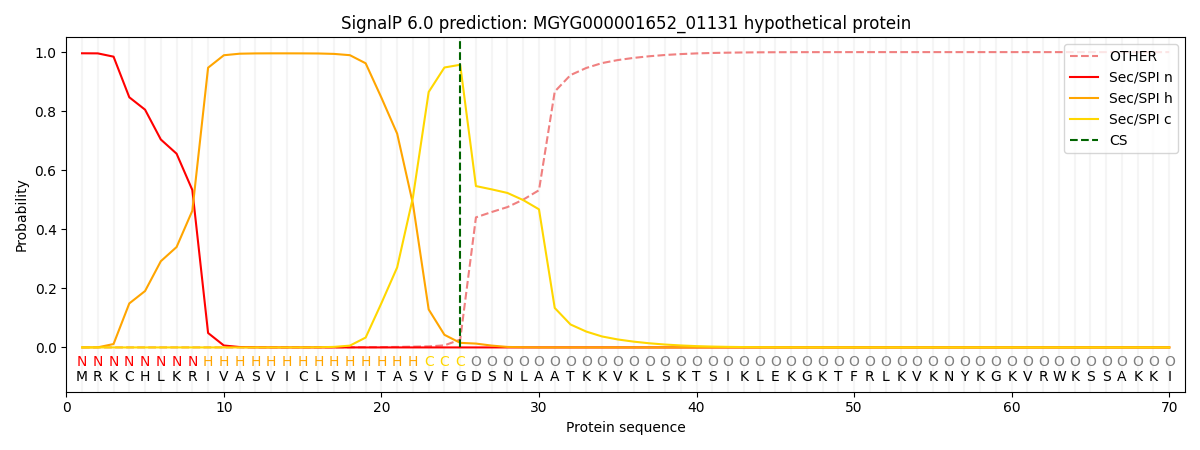
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000558 | 0.994635 | 0.004270 | 0.000203 | 0.000161 | 0.000143 |
