You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001654_02212
You are here: Home > Sequence: MGYG000001654_02212
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UMGS856 sp900760305 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Acutalibacteraceae; UMGS856; UMGS856 sp900760305 | |||||||||||
| CAZyme ID | MGYG000001654_02212 | |||||||||||
| CAZy Family | GT0 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1155; End: 2795 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd02513 | CMP-NeuAc_Synthase | 3.16e-74 | 2 | 217 | 1 | 223 | CMP-NeuAc_Synthase activates N-acetylneuraminic acid by adding CMP moiety. CMP-N-acetylneuraminic acid synthetase (CMP-NeuAc synthetase) or acylneuraminate cytidylyltransferase catalyzes the transfer the CMP moiety of CTP to the anomeric hydroxyl group of NeuAc in the presence of Mg++. It is the second to last step in the sialylation of the oligosaccharide component of glycoconjugates by providing the activated sugar-nucleotide cytidine 5'-monophosphate N-acetylneuraminic acid (CMP-Neu5Ac), the substrate for sialyltransferases. Eukaryotic CMP-NeuAc synthetases are predominantly located in the nucleus. The activated CMP-Neu5Ac diffuses from the nucleus into the cytoplasm. |
| COG1083 | NeuA | 1.06e-52 | 1 | 221 | 2 | 226 | CMP-N-acetylneuraminic acid synthetase [Cell wall/membrane/envelope biogenesis]. |
| COG3980 | SpsG | 1.92e-36 | 221 | 484 | 2 | 266 | Spore coat polysaccharide biosynthesis protein SpsG, predicted glycosyltransferase [Cell wall/membrane/envelope biogenesis]. |
| TIGR03590 | PseG | 3.18e-33 | 221 | 475 | 1 | 271 | UDP-2,4-diacetamido-2,4,6-trideoxy-beta-L-altropyranose hydrolase. This protein is found in association with enzymes involved in the biosynthesis of pseudaminic acid, a component of polysaccharide in certain Pseudomonas strains as well as a modification of flagellin in Campylobacter and Hellicobacter. The role of this protein is unclear, although it may participate in N-acetylation in conjunction with, or in the absence of PseH (TIGR03585) as it often scores above the trusted cutoff to pfam00583 representing a family of acetyltransferases. |
| pfam02348 | CTP_transf_3 | 2.92e-20 | 4 | 139 | 1 | 128 | Cytidylyltransferase. This family consists of two main Cytidylyltransferase activities: 1) 3-deoxy-manno-octulosonate cytidylyltransferase,, EC:2.7.7.38 catalyzing the reaction:- CTP + 3-deoxy-D-manno-octulosonate <=> diphosphate + CMP-3-deoxy-D-manno-octulosonate, 2) acylneuraminate cytidylyltransferase EC:2.7.7.43, catalyzing the reaction:- CTP + N-acylneuraminate <=> diphosphate + CMP-N-acylneuraminate. NeuAc cytydilyltransferase of Mannheimia haemolytica has been characterized describing kinetics and regulation by substrate charge, energetic charge and amino-sugar demand. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ACV55877.1 | 1.26e-184 | 3 | 546 | 7 | 546 |
| AMB92172.1 | 1.03e-169 | 1 | 544 | 1 | 543 |
| API89863.1 | 1.84e-167 | 1 | 545 | 1 | 543 |
| QGS37541.1 | 2.55e-162 | 1 | 546 | 6 | 553 |
| AMB95295.1 | 1.92e-159 | 1 | 544 | 1 | 543 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6IFD_A | 1.01e-26 | 4 | 222 | 23 | 249 | CrystalStructure of CMP-N-acetylneuraminate Synthetase from Vibrio cholerae in complex with CDP and Mg2+. [Vibrio cholerae],6IFD_B Crystal Structure of CMP-N-acetylneuraminate Synthetase from Vibrio cholerae in complex with CDP and Mg2+. [Vibrio cholerae],6IFD_C Crystal Structure of CMP-N-acetylneuraminate Synthetase from Vibrio cholerae in complex with CDP and Mg2+. [Vibrio cholerae],6IFD_D Crystal Structure of CMP-N-acetylneuraminate Synthetase from Vibrio cholerae in complex with CDP and Mg2+. [Vibrio cholerae],6IFI_A Crystal Structure of the Apo form of CMP-N-acetylneuraminate Synthetase from Vibrio cholerae [Vibrio cholerae],6IFI_B Crystal Structure of the Apo form of CMP-N-acetylneuraminate Synthetase from Vibrio cholerae [Vibrio cholerae] |
| 1QWJ_A | 7.43e-26 | 3 | 214 | 4 | 217 | TheCrystal Structure of Murine CMP-5-N-Acetylneuraminic Acid Synthetase [Mus musculus],1QWJ_B The Crystal Structure of Murine CMP-5-N-Acetylneuraminic Acid Synthetase [Mus musculus],1QWJ_C The Crystal Structure of Murine CMP-5-N-Acetylneuraminic Acid Synthetase [Mus musculus],1QWJ_D The Crystal Structure of Murine CMP-5-N-Acetylneuraminic Acid Synthetase [Mus musculus] |
| 6CKJ_A | 7.69e-16 | 4 | 221 | 6 | 227 | N.meningitidis CMP-sialic acid synthetase, ligand-free [Neisseria meningitidis],6CKK_A N. meningitidis CMP-sialic acid synthetase in the presence of CTP and Ca2+ [Neisseria meningitidis],6CKK_B N. meningitidis CMP-sialic acid synthetase in the presence of CTP and Ca2+ [Neisseria meningitidis],6CKL_A N. meningitidis CMP-sialic acid synthetase in the presence of CMP and Neu5Ac2en [Neisseria meningitidis],6CKL_B N. meningitidis CMP-sialic acid synthetase in the presence of CMP and Neu5Ac2en [Neisseria meningitidis],6CKL_C N. meningitidis CMP-sialic acid synthetase in the presence of CMP and Neu5Ac2en [Neisseria meningitidis],6CKM_A N. meningitidis CMP-sialic acid synthetase in the presence of CMP-sialic acid and Ca2+ [Neisseria meningitidis] |
| 4XWI_A | 5.90e-07 | 6 | 139 | 16 | 140 | X-rayCrystal structure of CMP-KDO Synthase from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],4XWI_B X-ray Crystal structure of CMP-KDO Synthase from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q58462 | 3.11e-28 | 221 | 479 | 2 | 275 | Uncharacterized protein MJ1062 OS=Methanocaldococcus jannaschii (strain ATCC 43067 / DSM 2661 / JAL-1 / JCM 10045 / NBRC 100440) OX=243232 GN=MJ1062 PE=1 SV=1 |
| Q8NFW8 | 3.41e-25 | 3 | 306 | 45 | 357 | N-acylneuraminate cytidylyltransferase OS=Homo sapiens OX=9606 GN=CMAS PE=1 SV=2 |
| Q5R6R5 | 3.41e-25 | 3 | 306 | 45 | 357 | N-acylneuraminate cytidylyltransferase OS=Pongo abelii OX=9601 GN=CMAS PE=2 SV=1 |
| P69060 | 8.22e-25 | 3 | 306 | 43 | 355 | N-acylneuraminate cytidylyltransferase OS=Rattus norvegicus OX=10116 GN=Cmas PE=2 SV=1 |
| Q99KK2 | 8.22e-25 | 3 | 306 | 43 | 355 | N-acylneuraminate cytidylyltransferase OS=Mus musculus OX=10090 GN=Cmas PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
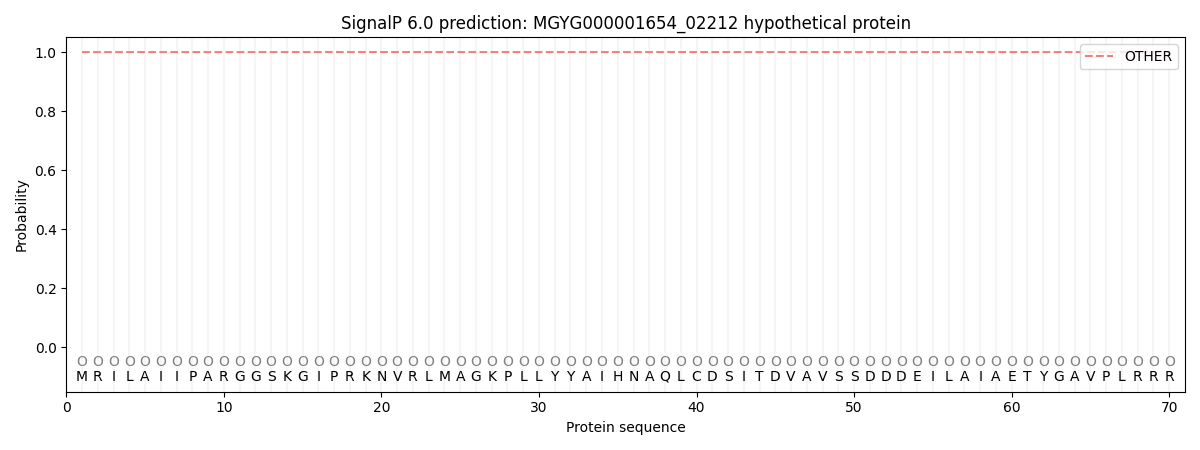
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000044 | 0.000018 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
