You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001659_02710
You are here: Home > Sequence: MGYG000001659_02710
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Paraprevotella sp900760455 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Paraprevotella; Paraprevotella sp900760455 | |||||||||||
| CAZyme ID | MGYG000001659_02710 | |||||||||||
| CAZy Family | CE1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 9662; End: 11908 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE1 | 505 | 716 | 6e-16 | 0.8678414096916299 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00930 | DPPIV_N | 1.08e-53 | 122 | 456 | 19 | 352 | Dipeptidyl peptidase IV (DPP IV) N-terminal region. This family is an alignment of the region to the N-terminal side of the active site. The Prosite motif does not correspond to this Pfam entry. |
| pfam00326 | Peptidase_S9 | 8.51e-46 | 539 | 734 | 2 | 210 | Prolyl oligopeptidase family. |
| COG1506 | DAP2 | 2.57e-37 | 167 | 715 | 68 | 594 | Dipeptidyl aminopeptidase/acylaminoacyl peptidase [Amino acid transport and metabolism]. |
| COG0412 | DLH | 4.66e-07 | 547 | 728 | 50 | 215 | Dienelactone hydrolase [Secondary metabolites biosynthesis, transport and catabolism]. |
| pfam00756 | Esterase | 5.70e-06 | 499 | 729 | 5 | 235 | Putative esterase. This family contains Esterase D. However it is not clear if all members of the family have the same function. This family is related to the pfam00135 family. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AJY87077.1 | 1.71e-201 | 69 | 743 | 269 | 950 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2D5L_A | 5.21e-55 | 167 | 720 | 129 | 690 | CrystalStructure of Prolyl Tripeptidyl Aminopeptidase from Porphyromonas gingivalis [Porphyromonas gingivalis W83],2EEP_A Prolyl Tripeptidyl Aminopeptidase Complexed with an Inhibitor [Porphyromonas gingivalis W83] |
| 2Z3W_A | 9.71e-55 | 167 | 720 | 129 | 690 | ChainA, Dipeptidyl aminopeptidase IV [Porphyromonas gingivalis W83],2Z3Z_A Chain A, Dipeptidyl aminopeptidase IV [Porphyromonas gingivalis W83] |
| 2DCM_A | 1.33e-54 | 167 | 720 | 129 | 690 | ChainA, dipeptidyl aminopeptidase IV, putative [Porphyromonas gingivalis W83] |
| 2ECF_A | 4.95e-48 | 47 | 725 | 40 | 727 | CrystalStructure of Dipeptidyl Aminopeptidase IV from Stenotrophomonas maltophilia [Stenotrophomonas maltophilia] |
| 5YP1_A | 1.91e-46 | 117 | 734 | 122 | 743 | Crystalstructure of dipeptidyl peptidase IV (DPP IV) from Pseudoxanthomonas mexicana WO24 [Pseudoxanthomonas mexicana],5YP1_B Crystal structure of dipeptidyl peptidase IV (DPP IV) from Pseudoxanthomonas mexicana WO24 [Pseudoxanthomonas mexicana],5YP1_C Crystal structure of dipeptidyl peptidase IV (DPP IV) from Pseudoxanthomonas mexicana WO24 [Pseudoxanthomonas mexicana],5YP1_D Crystal structure of dipeptidyl peptidase IV (DPP IV) from Pseudoxanthomonas mexicana WO24 [Pseudoxanthomonas mexicana],5YP2_A Crystal structure of dipeptidyl peptidase IV (DPP IV) with DPP4 inhibitor from Pseudoxanthomonas mexicana WO24 [Pseudoxanthomonas mexicana],5YP2_B Crystal structure of dipeptidyl peptidase IV (DPP IV) with DPP4 inhibitor from Pseudoxanthomonas mexicana WO24 [Pseudoxanthomonas mexicana],5YP3_A Crystal structure of dipeptidyl peptidase IV (DPP IV) with Ile-Pro from Pseudoxanthomonas mexicana [Pseudoxanthomonas mexicana],5YP3_B Crystal structure of dipeptidyl peptidase IV (DPP IV) with Ile-Pro from Pseudoxanthomonas mexicana [Pseudoxanthomonas mexicana],5YP3_C Crystal structure of dipeptidyl peptidase IV (DPP IV) with Ile-Pro from Pseudoxanthomonas mexicana [Pseudoxanthomonas mexicana],5YP3_D Crystal structure of dipeptidyl peptidase IV (DPP IV) with Ile-Pro from Pseudoxanthomonas mexicana [Pseudoxanthomonas mexicana],5YP4_A Crystal structure of dipeptidyl peptidase IV (DPP IV) with Lys-Pro from Pseudoxanthomonas mexicana WO24 [Pseudoxanthomonas mexicana],5YP4_B Crystal structure of dipeptidyl peptidase IV (DPP IV) with Lys-Pro from Pseudoxanthomonas mexicana WO24 [Pseudoxanthomonas mexicana],5YP4_C Crystal structure of dipeptidyl peptidase IV (DPP IV) with Lys-Pro from Pseudoxanthomonas mexicana WO24 [Pseudoxanthomonas mexicana],5YP4_D Crystal structure of dipeptidyl peptidase IV (DPP IV) with Lys-Pro from Pseudoxanthomonas mexicana WO24 [Pseudoxanthomonas mexicana] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q7MUW6 | 3.99e-54 | 167 | 720 | 155 | 716 | Prolyl tripeptidyl peptidase OS=Porphyromonas gingivalis (strain ATCC BAA-308 / W83) OX=242619 GN=ptpA PE=1 SV=1 |
| B2RJX3 | 1.01e-53 | 167 | 720 | 155 | 716 | Prolyl tripeptidyl peptidase OS=Porphyromonas gingivalis (strain ATCC 33277 / DSM 20709 / CIP 103683 / JCM 12257 / NCTC 11834 / 2561) OX=431947 GN=ptpA PE=3 SV=1 |
| Q6F3I7 | 1.05e-45 | 117 | 734 | 122 | 743 | Dipeptidyl aminopeptidase 4 OS=Pseudoxanthomonas mexicana OX=128785 GN=dap4 PE=1 SV=1 |
| P97321 | 1.64e-30 | 49 | 734 | 59 | 753 | Prolyl endopeptidase FAP OS=Mus musculus OX=10090 GN=Fap PE=1 SV=1 |
| Q5B934 | 2.96e-30 | 167 | 717 | 268 | 855 | Probable dipeptidyl-aminopeptidase B OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=dapB PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
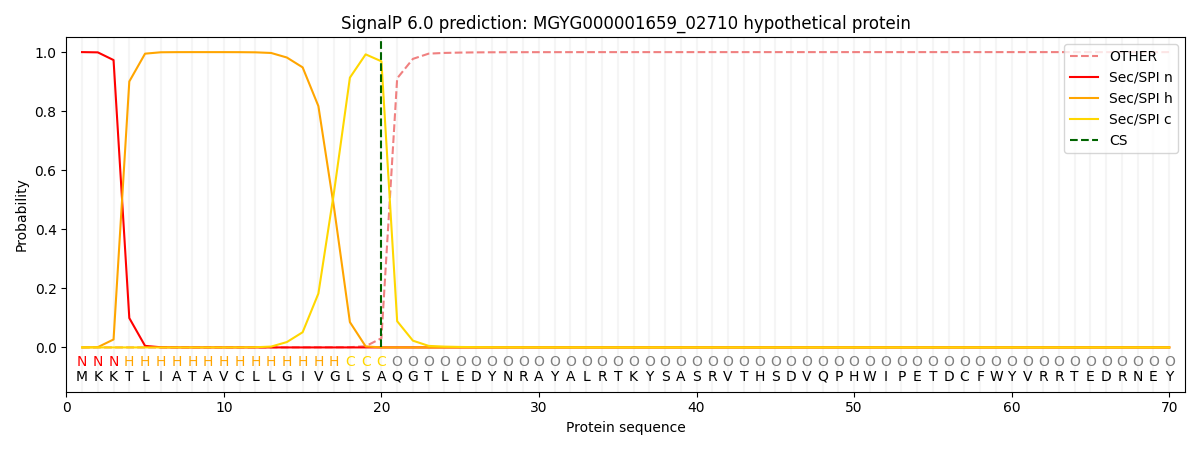
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000842 | 0.998176 | 0.000387 | 0.000195 | 0.000182 | 0.000186 |

