You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001666_01802
Basic Information
help
| Species |
|
| Lineage |
Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Muribaculaceae; CAG-873;
|
| CAZyme ID |
MGYG000001666_01802
|
| CAZy Family |
GH5 |
| CAZyme Description |
hypothetical protein
|
| CAZyme Property |
| Protein Length |
CGC |
Molecular Weight |
Isoelectric Point |
| 673 |
|
74053.44 |
5.1371 |
|
| Genome Property |
| Genome Assembly ID |
Genome Size |
Genome Type |
Country |
Continent |
| MGYG000001666 |
2832137 |
MAG |
United States |
North America |
|
| Gene Location |
Start: 6491;
End: 8512
Strand: -
|
| Family |
Start |
End |
Evalue |
family coverage |
| GH5 |
38 |
315 |
1.1e-115 |
0.9884169884169884 |
| Cdd ID |
Domain |
E-Value |
qStart |
qEnd |
sStart |
sEnd |
Domain Description |
| pfam00150
|
Cellulase |
1.01e-14 |
32 |
266 |
2 |
201 |
Cellulase (glycosyl hydrolase family 5). |
This protein is predicted as SP
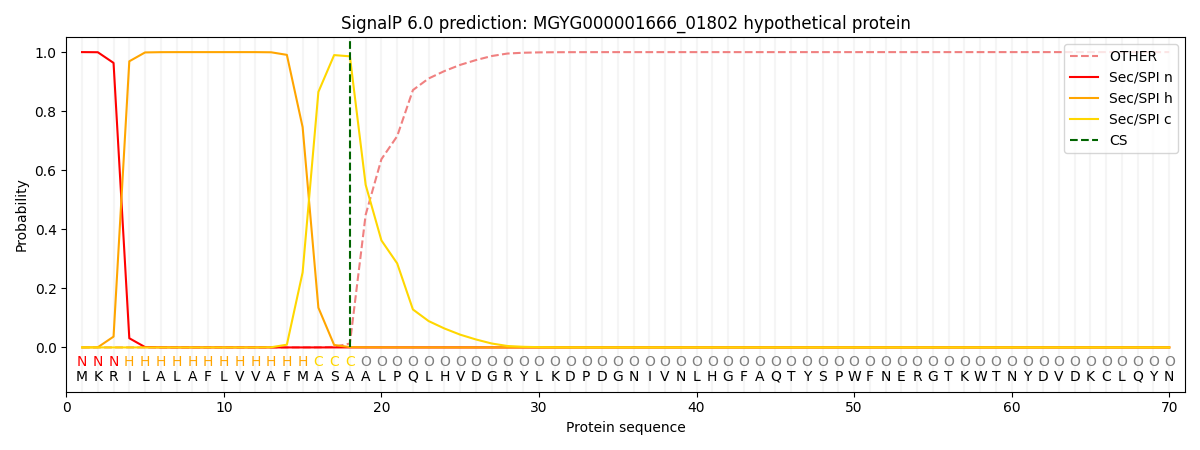
| Other |
SP_Sec_SPI |
LIPO_Sec_SPII |
TAT_Tat_SPI |
TATLIP_Sec_SPII |
PILIN_Sec_SPIII |
|
0.000226
|
0.999185
|
0.000143
|
0.000141
|
0.000129
|
0.000128
|
There is no transmembrane helices in MGYG000001666_01802.

