You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001681_01486
You are here: Home > Sequence: MGYG000001681_01486
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Ruminococcus sp900761275 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Ruminococcaceae; Ruminococcus; Ruminococcus sp900761275 | |||||||||||
| CAZyme ID | MGYG000001681_01486 | |||||||||||
| CAZy Family | GH9 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 122; End: 2374 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH9 | 33 | 452 | 1e-97 | 0.9976076555023924 |
| CBM79 | 526 | 633 | 9.3e-40 | 0.9545454545454546 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00759 | Glyco_hydro_9 | 1.15e-93 | 36 | 451 | 1 | 374 | Glycosyl hydrolase family 9. |
| PLN02613 | PLN02613 | 1.48e-45 | 4 | 470 | 1 | 498 | endoglucanase |
| PLN02345 | PLN02345 | 4.15e-45 | 38 | 467 | 2 | 468 | endoglucanase |
| PLN00119 | PLN00119 | 5.96e-39 | 33 | 454 | 31 | 488 | endoglucanase |
| PLN02420 | PLN02420 | 2.56e-38 | 6 | 454 | 14 | 506 | endoglucanase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| EWM53237.1 | 8.34e-185 | 2 | 746 | 11 | 880 |
| CBL17554.1 | 1.38e-180 | 4 | 471 | 17 | 465 |
| CDE33541.1 | 1.16e-174 | 2 | 578 | 20 | 587 |
| AEY66060.1 | 6.15e-103 | 9 | 455 | 20 | 514 |
| AEV68415.1 | 1.04e-99 | 1 | 458 | 14 | 518 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2YIK_A | 3.62e-99 | 6 | 462 | 12 | 520 | ChainA, Endoglucanase [Acetivibrio thermocellus] |
| 1IA6_A | 5.70e-76 | 30 | 453 | 2 | 425 | CrystalStructure Of The Cellulase Cel9m Of C. Cellulolyticum [Ruminiclostridium cellulolyticum],1IA7_A Crystal Structure Of The Cellulase Cel9m Of C. Cellulolyticium In Complex With Cellobiose [Ruminiclostridium cellulolyticum] |
| 2XFG_A | 1.70e-65 | 28 | 458 | 20 | 463 | ChainA, ENDOGLUCANASE 1 [Acetivibrio thermocellus] |
| 4DOD_A | 6.98e-60 | 28 | 458 | 22 | 463 | Thestructure of Cbescii CelA GH9 module [Caldicellulosiruptor bescii],4DOE_A The liganded structure of Cbescii CelA GH9 module [Caldicellulosiruptor bescii] |
| 1K72_A | 6.03e-58 | 30 | 480 | 2 | 461 | TheX-ray Crystal Structure Of Cel9G Complexed With cellotriose [Ruminiclostridium cellulolyticum],1K72_B The X-ray Crystal Structure Of Cel9G Complexed With cellotriose [Ruminiclostridium cellulolyticum] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q02934 | 7.41e-62 | 28 | 473 | 72 | 529 | Endoglucanase 1 OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celI PE=1 SV=2 |
| P37700 | 5.35e-57 | 28 | 480 | 35 | 496 | Endoglucanase G OS=Ruminiclostridium cellulolyticum (strain ATCC 35319 / DSM 5812 / JCM 6584 / H10) OX=394503 GN=celCCG PE=1 SV=2 |
| P22534 | 4.03e-54 | 26 | 458 | 20 | 463 | Endoglucanase A OS=Caldicellulosiruptor saccharolyticus OX=44001 GN=celA PE=3 SV=2 |
| P26225 | 1.09e-50 | 2 | 458 | 9 | 478 | Endoglucanase B OS=Cellulomonas fimi OX=1708 GN=cenB PE=3 SV=1 |
| P26224 | 6.21e-50 | 7 | 458 | 11 | 467 | Endoglucanase F OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celF PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
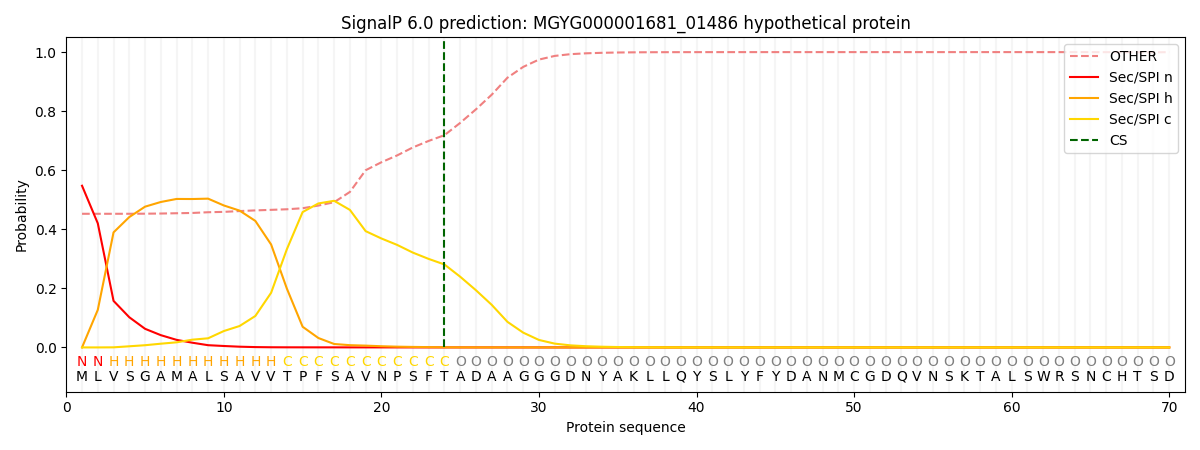
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.461556 | 0.537576 | 0.000190 | 0.000297 | 0.000193 | 0.000187 |
