You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001696_01127
You are here: Home > Sequence: MGYG000001696_01127
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
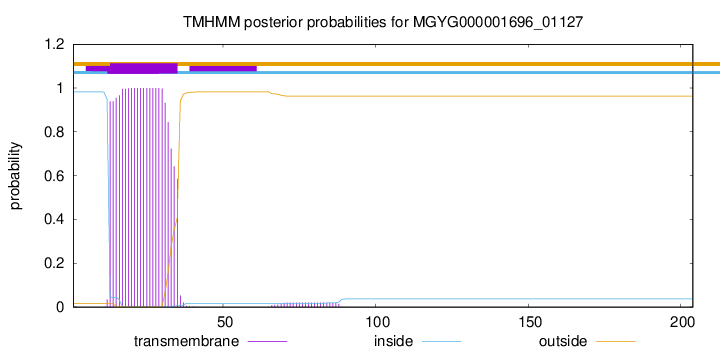
TMHMM annotations
Basic Information help
| Species | Enterococcus_A raffinosus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Enterococcaceae; Enterococcus_A; Enterococcus_A raffinosus | |||||||||||
| CAZyme ID | MGYG000001696_01127 | |||||||||||
| CAZy Family | GH73 | |||||||||||
| CAZyme Description | Exo-glucosaminidase LytG | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 18778; End: 19392 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1705 | FlgJ | 2.49e-73 | 17 | 202 | 11 | 188 | Flagellum-specific peptidoglycan hydrolase FlgJ [Cell wall/membrane/envelope biogenesis, Cell motility]. |
| PRK08581 | PRK08581 | 2.06e-39 | 37 | 204 | 310 | 474 | amidase domain-containing protein. |
| smart00047 | LYZ2 | 2.67e-39 | 53 | 202 | 9 | 147 | Lysozyme subfamily 2. Eubacterial enzymes distantly related to eukaryotic lysozymes. |
| PRK05684 | flgJ | 5.45e-38 | 51 | 195 | 151 | 296 | flagellar assembly peptidoglycan hydrolase FlgJ. |
| TIGR02541 | flagell_FlgJ | 1.10e-36 | 53 | 193 | 149 | 290 | flagellar rod assembly protein/muramidase FlgJ. The N-terminal region of this protein acts directly in flagellar rod assembly. The C-terminal region is a flagellum-specific muramidase (peptidoglycan hydrolase) required for formation of the outer membrane L ring. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QZO10089.1 | 1.03e-145 | 1 | 204 | 1 | 204 |
| QXJ60493.1 | 5.97e-145 | 1 | 204 | 1 | 204 |
| BBM18487.1 | 7.20e-137 | 1 | 204 | 1 | 204 |
| AYQ23607.1 | 4.47e-131 | 1 | 204 | 1 | 204 |
| QCQ13336.1 | 4.47e-131 | 1 | 204 | 1 | 204 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3FI7_A | 1.08e-30 | 30 | 202 | 9 | 183 | CrystalStructure of the autolysin Auto (Lmo1076) from Listeria monocytogenes, catalytic domain [Listeria monocytogenes EGD-e] |
| 5DN5_A | 4.21e-24 | 50 | 194 | 1 | 146 | Structureof a C-terminally truncated glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],5DN5_B Structure of a C-terminally truncated glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],5DN5_C Structure of a C-terminally truncated glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2] |
| 5DN4_A | 6.14e-24 | 50 | 194 | 1 | 146 | Structureof the glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2] |
| 3VWO_A | 4.27e-20 | 53 | 200 | 2 | 150 | Crystalstructure of peptidoglycan hydrolase mutant from Sphingomonas sp. A1 [Sphingomonas sp. A1] |
| 2ZYC_A | 5.27e-20 | 53 | 200 | 3 | 151 | ChainA, Peptidoglycan hydrolase FlgJ [Sphingomonas sp. A1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O32083 | 2.82e-48 | 7 | 202 | 5 | 198 | Exo-glucosaminidase LytG OS=Bacillus subtilis (strain 168) OX=224308 GN=lytG PE=1 SV=1 |
| P0C2T5 | 8.49e-27 | 10 | 202 | 14 | 214 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. cremoris OX=1359 GN=acmA PE=3 SV=1 |
| Q9CIT4 | 1.65e-26 | 19 | 202 | 30 | 214 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. lactis (strain IL1403) OX=272623 GN=acmA PE=3 SV=1 |
| A2RHZ5 | 3.10e-26 | 8 | 202 | 14 | 214 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. cremoris (strain MG1363) OX=416870 GN=acmA PE=3 SV=1 |
| P58231 | 2.48e-24 | 55 | 194 | 152 | 292 | Peptidoglycan hydrolase FlgJ OS=Escherichia coli O157:H7 OX=83334 GN=flgJ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
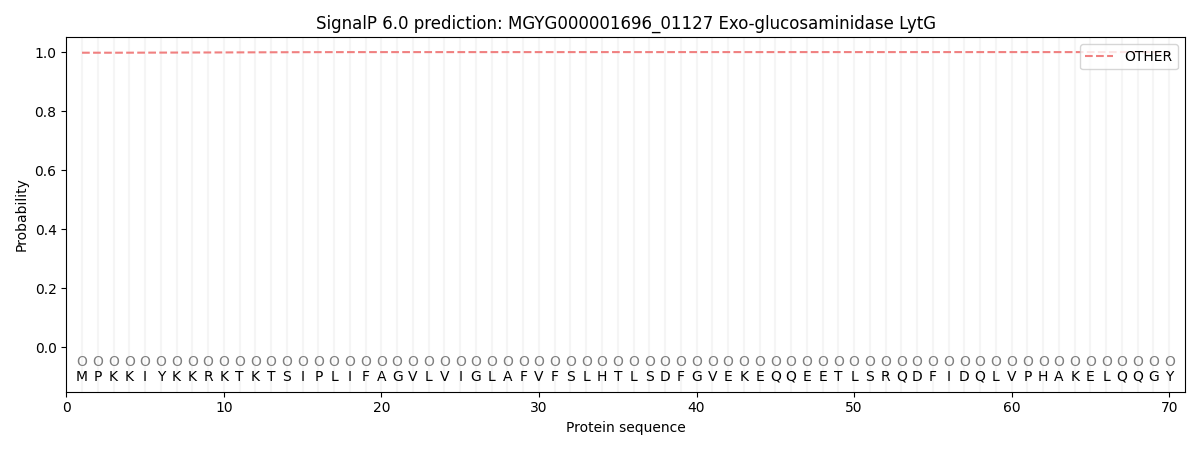
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.997964 | 0.000258 | 0.000020 | 0.000001 | 0.000001 | 0.001789 |