You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001702_01789
You are here: Home > Sequence: MGYG000001702_01789
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
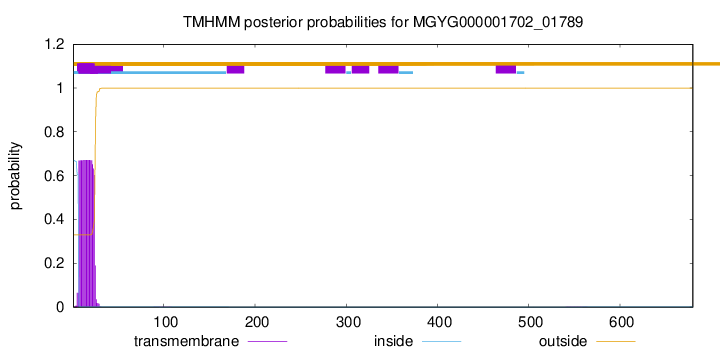
TMHMM annotations
Basic Information help
| Species | Virgibacillus picturae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Bacillales_D; Amphibacillaceae; Virgibacillus; Virgibacillus picturae | |||||||||||
| CAZyme ID | MGYG000001702_01789 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | Beta-hexosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1718592; End: 1720634 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 129 | 354 | 4.6e-67 | 0.9768518518518519 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1472 | BglX | 5.35e-108 | 48 | 468 | 1 | 381 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| pfam00933 | Glyco_hydro_3 | 8.12e-101 | 49 | 390 | 1 | 316 | Glycosyl hydrolase family 3 N terminal domain. |
| PRK15098 | PRK15098 | 2.60e-50 | 29 | 575 | 22 | 600 | beta-glucosidase BglX. |
| PRK05337 | PRK05337 | 2.88e-47 | 95 | 359 | 26 | 285 | beta-hexosaminidase; Provisional |
| pfam01915 | Glyco_hydro_3_C | 3.80e-22 | 427 | 586 | 1 | 216 | Glycosyl hydrolase family 3 C-terminal domain. This domain is involved in catalysis and may be involved in binding beta-glucan. This domain is found associated with pfam00933. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AVR01146.1 | 0.0 | 25 | 680 | 1 | 656 |
| QRZ20287.1 | 0.0 | 33 | 679 | 13 | 658 |
| BAC13255.1 | 0.0 | 9 | 676 | 9 | 666 |
| AQS56346.1 | 1.72e-296 | 19 | 676 | 2 | 658 |
| ASK62321.1 | 1.21e-294 | 1 | 677 | 1 | 677 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6K5J_A | 1.21e-107 | 48 | 574 | 11 | 531 | Structureof a glycoside hydrolase family 3 beta-N-acetylglucosaminidase from Paenibacillus sp. str. FPU-7 [Paenibacillaceae] |
| 3BMX_A | 4.10e-98 | 23 | 584 | 20 | 627 | Beta-N-hexosaminidase(YbbD) from Bacillus subtilis [Bacillus subtilis],3BMX_B Beta-N-hexosaminidase (YbbD) from Bacillus subtilis [Bacillus subtilis],3NVD_A Structure of YBBD in complex with pugnac [Bacillus subtilis],3NVD_B Structure of YBBD in complex with pugnac [Bacillus subtilis] |
| 4GYJ_A | 2.58e-97 | 23 | 584 | 24 | 631 | Crystalstructure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1) [Bacillus subtilis subsp. subtilis str. 168],4GYJ_B Crystal structure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1) [Bacillus subtilis subsp. subtilis str. 168],4GYK_A Crystal structure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1211) [Bacillus subtilis subsp. subtilis str. 168],4GYK_B Crystal structure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1211) [Bacillus subtilis subsp. subtilis str. 168] |
| 3LK6_A | 3.13e-97 | 40 | 584 | 8 | 601 | ChainA, Lipoprotein ybbD [Bacillus subtilis],3LK6_B Chain B, Lipoprotein ybbD [Bacillus subtilis],3LK6_C Chain C, Lipoprotein ybbD [Bacillus subtilis],3LK6_D Chain D, Lipoprotein ybbD [Bacillus subtilis] |
| 4ZM6_A | 1.15e-85 | 46 | 577 | 5 | 530 | Aunique GCN5-related glucosamine N-acetyltransferase region exist in the fungal multi-domain GH3 beta-N-acetylglucosaminidase [Rhizomucor miehei CAU432],4ZM6_B A unique GCN5-related glucosamine N-acetyltransferase region exist in the fungal multi-domain GH3 beta-N-acetylglucosaminidase [Rhizomucor miehei CAU432] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P40406 | 2.24e-97 | 23 | 584 | 20 | 627 | Beta-hexosaminidase OS=Bacillus subtilis (strain 168) OX=224308 GN=nagZ PE=1 SV=1 |
| P48823 | 3.02e-83 | 88 | 574 | 57 | 596 | Beta-hexosaminidase A OS=Pseudoalteromonas piscicida OX=43662 GN=cht60 PE=1 SV=1 |
| Q7WUL3 | 9.96e-50 | 41 | 572 | 19 | 553 | Beta-N-acetylglucosaminidase/beta-glucosidase OS=Cellulomonas fimi OX=1708 GN=nag3 PE=1 SV=1 |
| A7LXU3 | 2.36e-41 | 38 | 594 | 36 | 636 | Beta-glucosidase BoGH3B OS=Bacteroides ovatus (strain ATCC 8483 / DSM 1896 / JCM 5824 / BCRC 10623 / CCUG 4943 / NCTC 11153) OX=411476 GN=BACOVA_02659 PE=1 SV=1 |
| Q3SKU2 | 1.59e-36 | 95 | 333 | 28 | 259 | Beta-hexosaminidase OS=Thiobacillus denitrificans (strain ATCC 25259) OX=292415 GN=nagZ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
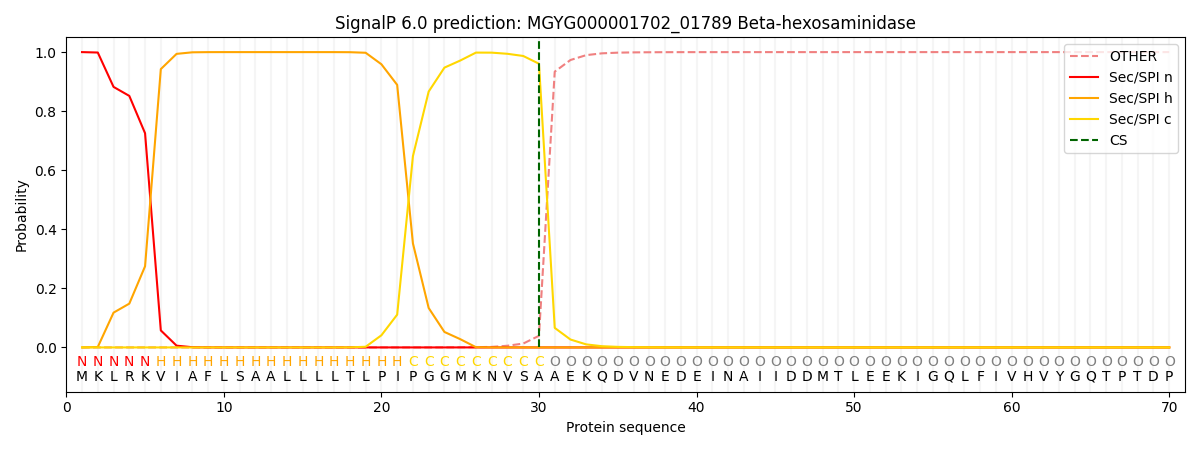
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000258 | 0.999023 | 0.000238 | 0.000161 | 0.000152 | 0.000143 |