You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001725_01333
You are here: Home > Sequence: MGYG000001725_01333
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Massilimaliae sp900752435 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Ruminococcaceae; Massilimaliae; Massilimaliae sp900752435 | |||||||||||
| CAZyme ID | MGYG000001725_01333 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | D-alanine--poly(phosphoribitol) ligase subunit 1 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 6505; End: 14016 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3321 | PksD | 0.0 | 14 | 862 | 6 | 862 | Acyl transferase domain in polyketide synthase (PKS) enzymes [Secondary metabolites biosynthesis, transport and catabolism]. |
| cd00833 | PKS | 7.07e-180 | 13 | 431 | 2 | 421 | polyketide synthases (PKSs) polymerize simple fatty acids into a large variety of different products, called polyketides, by successive decarboxylating Claisen condensations. PKSs can be divided into 2 groups, modular type I PKSs consisting of one or more large multifunctional proteins and iterative type II PKSs, complexes of several monofunctional subunits. |
| cd12114 | A_NRPS_TlmIV_like | 2.93e-174 | 1452 | 1926 | 2 | 477 | The adenylation domain of nonribosomal peptide synthetases (NRPS), including Streptoalloteichus tallysomycin biosynthesis genes. The adenylation (A) domain of NRPS recognizes a specific amino acid or hydroxy acid and activates it as an (amino) acyl adenylate by hydrolysis of ATP. The activated acyl moiety then forms a thioester to the enzyme-bound cofactor phosphopantetheine of a peptidyl carrier protein domain. NRPSs are large multifunctional enzymes which synthesize many therapeutically useful peptides in bacteria and fungi via a template-directed, nucleic acid independent nonribosomal mechanism. These natural products include antibiotics, immunosuppressants, plant and animal toxins, and enzyme inhibitors. NRPS has a distinct modular structure in which each module is responsible for the recognition, activation, and in some cases, modification of a single amino acid residue of the final peptide product. The modules can be subdivided into domains that catalyze specific biochemical reactions. This family includes the TLM biosynthetic gene cluster from Streptoalloteichus that consists of nine NRPS genes; the N-terminal module of TlmVI (NRPS-5) and the starter module of BlmVI (NRPS-5) are comprised of the acyl CoA ligase (AL) and acyl carrier protein (ACP)-like domains, which are thought to be involved in the biosynthesis of the beta-aminoalaninamide moiety. |
| cd05930 | A_NRPS | 3.93e-136 | 1451 | 1926 | 1 | 444 | The adenylation domain of nonribosomal peptide synthetases (NRPS). The adenylation (A) domain of NRPS recognizes a specific amino acid or hydroxy acid and activates it as an (amino) acyl adenylate by hydrolysis of ATP. The activated acyl moiety then forms a thioester bond to the enzyme-bound cofactor phosphopantetheine of a peptidyl carrier protein domain. NRPSs are large multifunctional enzymes which synthesize many therapeutically useful peptides in bacteria and fungi via a template-directed, nucleic acid independent nonribosomal mechanism. These natural products include antibiotics, immunosuppressants, plant and animal toxins, and enzyme inhibitors. NRPS has a distinct modular structure in which each module is responsible for the recognition, activation, and in some cases, modification of a single amino acid residue of the final peptide product. The modules can be subdivided into domains that catalyze specific biochemical reactions. |
| COG1020 | EntF | 3.79e-132 | 1208 | 1845 | 1 | 634 | Non-ribosomal peptide synthetase component F [Secondary metabolites biosynthesis, transport and catabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BAZ00088.1 | 4.99e-195 | 13 | 2010 | 1213 | 3289 |
| BAZ75991.1 | 4.99e-195 | 13 | 2010 | 1213 | 3289 |
| BAY30132.1 | 1.60e-193 | 13 | 2010 | 1215 | 3291 |
| BAY90071.1 | 6.71e-190 | 13 | 2010 | 1212 | 3280 |
| AFY93865.1 | 1.13e-112 | 13 | 956 | 960 | 1968 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4MZ0_A | 1.58e-160 | 14 | 869 | 41 | 933 | ChainA, CurL [Moorena producens 3L],4MZ0_B Chain B, CurL [Moorena producens 3L] |
| 7LY7_A | 2.74e-160 | 992 | 1984 | 11 | 988 | ChainA, BmdB, Bacillamide NRPS [Thermoactinomyces vulgaris] |
| 7LY4_E | 2.81e-160 | 992 | 1984 | 11 | 988 | ChainE, BmdB, bacillamide NRPS [Thermoactinomyces vulgaris] |
| 7VEE_A | 5.73e-146 | 14 | 865 | 23 | 905 | ChainA, Polyketide synthase [Streptomyces graminofaciens],7VEF_A Chain A, Polyketide synthase [Streptomyces graminofaciens] |
| 6LTA_A | 1.32e-143 | 989 | 2010 | 54 | 1095 | ChainA, Nonribosomal peptide synthetase [Streptomyces sp. Sp080513GE-23],6LTB_A Chain A, Nonribosomal peptide synthetase [Streptomyces sp. Sp080513GE-23],6LTC_A Chain A, Nonribosomal peptide synthetase [Streptomyces sp. Sp080513GE-23],6LTD_A Chain A, Nonribosomal peptide synthetase [Streptomyces sp. Sp080513GE-23],6LTD_B Chain B, Nonribosomal peptide synthetase [Streptomyces sp. Sp080513GE-23] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P48633 | 3.00e-175 | 900 | 2498 | 24 | 1900 | High-molecular-weight protein 2 OS=Yersinia enterocolitica serotype O:8 / biotype 1B (strain NCTC 13174 / 8081) OX=393305 GN=irp2 PE=3 SV=1 |
| O68006 | 1.87e-155 | 910 | 2415 | 556 | 2000 | Bacitracin synthase 1 OS=Bacillus licheniformis OX=1402 GN=bacA PE=3 SV=1 |
| Q7TYQ4 | 3.47e-137 | 897 | 1936 | 8 | 1047 | Phenyloxazoline synthase MbtB OS=Mycobacterium bovis (strain ATCC BAA-935 / AF2122/97) OX=233413 GN=mbtB PE=3 SV=1 |
| P9WQ63 | 4.67e-137 | 897 | 1936 | 8 | 1047 | Phenyloxazoline synthase MbtB OS=Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) OX=83332 GN=mbtB PE=1 SV=1 |
| P9WQ62 | 1.14e-136 | 897 | 1936 | 8 | 1047 | Phenyloxazoline synthase MbtB OS=Mycobacterium tuberculosis (strain CDC 1551 / Oshkosh) OX=83331 GN=mbtB PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
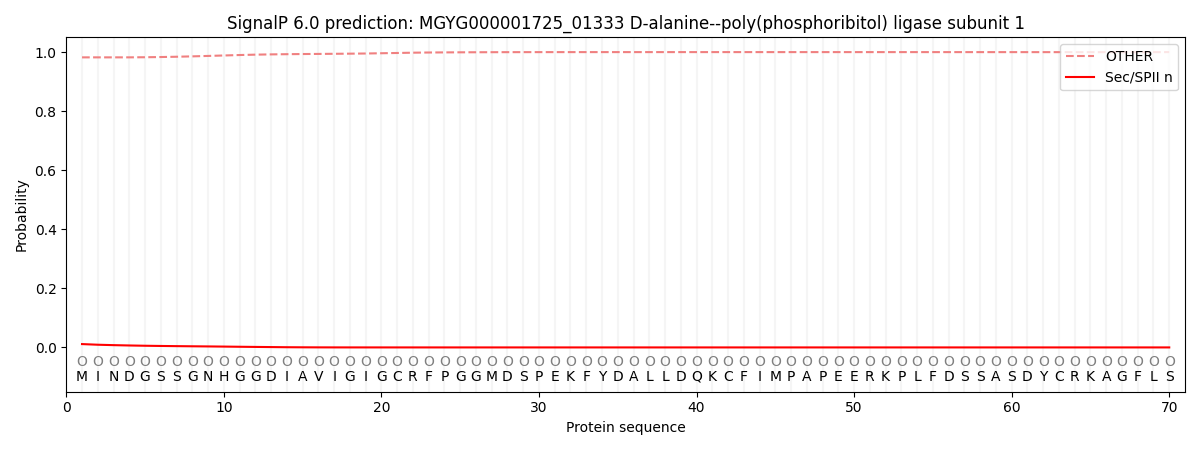
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.982763 | 0.005625 | 0.011402 | 0.000045 | 0.000017 | 0.000153 |
