You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001750_02287
You are here: Home > Sequence: MGYG000001750_02287
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phocaeicola ilei | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Phocaeicola; Phocaeicola ilei | |||||||||||
| CAZyme ID | MGYG000001750_02287 | |||||||||||
| CAZy Family | GH28 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 5577; End: 7028 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH28 | 53 | 399 | 9.6e-66 | 0.9230769230769231 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG5434 | Pgu1 | 1.72e-66 | 26 | 463 | 82 | 509 | Polygalacturonase [Carbohydrate transport and metabolism]. |
| PLN02218 | PLN02218 | 6.09e-10 | 28 | 336 | 69 | 350 | polygalacturonase ADPG |
| pfam12708 | Pectate_lyase_3 | 1.86e-08 | 28 | 82 | 3 | 59 | Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28. |
| pfam00295 | Glyco_hydro_28 | 2.44e-07 | 114 | 368 | 36 | 273 | Glycosyl hydrolases family 28. Glycosyl hydrolase family 28 includes polygalacturonase EC:3.2.1.15 as well as rhamnogalacturonase A(RGase A), EC:3.2.1.-. These enzymes are important in cell wall metabolism. |
| PLN02188 | PLN02188 | 3.20e-05 | 3 | 250 | 7 | 229 | polygalacturonase/glycoside hydrolase family protein |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| VTZ55482.1 | 2.33e-274 | 1 | 475 | 1 | 471 |
| ADY37530.1 | 2.68e-270 | 11 | 474 | 8 | 470 |
| QQY43062.1 | 1.24e-263 | 23 | 476 | 17 | 469 |
| QUT56815.1 | 3.55e-263 | 23 | 476 | 17 | 469 |
| ABR37914.1 | 3.55e-263 | 23 | 476 | 17 | 469 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3JUR_A | 5.49e-19 | 28 | 391 | 29 | 407 | Thecrystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_B The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_C The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_D The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima] |
| 4MXN_A | 3.70e-13 | 23 | 221 | 18 | 215 | Crystalstructure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_B Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_C Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_D Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184] |
| 2PYG_A | 2.16e-08 | 27 | 92 | 3 | 68 | Azotobactervinelandii Mannuronan C-5 epimerase AlgE4 A-module [Azotobacter vinelandii],2PYG_B Azotobacter vinelandii Mannuronan C-5 epimerase AlgE4 A-module [Azotobacter vinelandii],2PYH_A Azotobacter vinelandii Mannuronan C-5 epimerase AlgE4 A-module complexed with mannuronan trisaccharide [Azotobacter vinelandii],2PYH_B Azotobacter vinelandii Mannuronan C-5 epimerase AlgE4 A-module complexed with mannuronan trisaccharide [Azotobacter vinelandii] |
| 3ZPP_A | 2.58e-08 | 28 | 263 | 25 | 256 | Structureof the Streptococcus pneumoniae surface protein and adhesin PfbA [Streptococcus pneumoniae TIGR4] |
| 4MR0_A | 3.34e-08 | 28 | 263 | 117 | 348 | Crystalstructure of PfbA, a surface adhesin of Streptococcus pneumoniae [Streptococcus pneumoniae R6],4MR0_B Crystal structure of PfbA, a surface adhesin of Streptococcus pneumoniae [Streptococcus pneumoniae R6] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A7PZL3 | 1.51e-13 | 21 | 329 | 57 | 359 | Probable polygalacturonase OS=Vitis vinifera OX=29760 GN=GSVIVT00026920001 PE=1 SV=1 |
| P15922 | 1.15e-09 | 25 | 340 | 150 | 492 | Exo-poly-alpha-D-galacturonosidase OS=Dickeya chrysanthemi OX=556 GN=pehX PE=1 SV=1 |
| Q7M1E7 | 3.14e-09 | 24 | 290 | 56 | 310 | Polygalacturonase OS=Chamaecyparis obtusa OX=13415 PE=1 SV=1 |
| Q9ZFG9 | 3.18e-09 | 27 | 92 | 3 | 68 | Alginate lyase 7 OS=Azotobacter vinelandii OX=354 GN=algE7 PE=1 SV=1 |
| Q44494 | 4.40e-08 | 27 | 92 | 3 | 68 | Mannuronan C5-epimerase AlgE1 OS=Azotobacter vinelandii OX=354 GN=algE1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
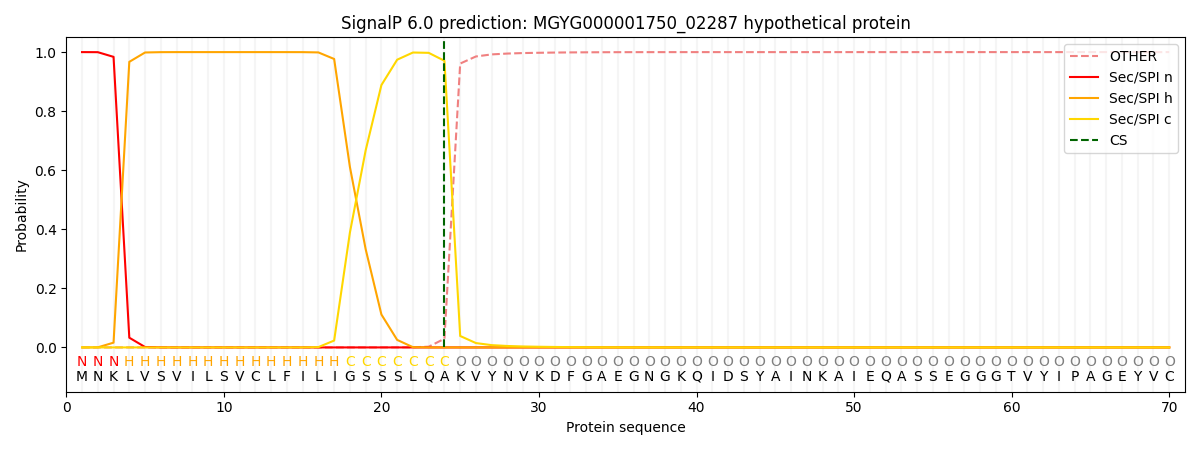
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000329 | 0.998889 | 0.000203 | 0.000188 | 0.000179 | 0.000175 |
