You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001770_00836
You are here: Home > Sequence: MGYG000001770_00836
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp900313215 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900313215 | |||||||||||
| CAZyme ID | MGYG000001770_00836 | |||||||||||
| CAZy Family | PL1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 34911; End: 38411 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL1 | 76 | 254 | 1.5e-49 | 0.9890710382513661 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd00063 | FN3 | 1.75e-12 | 435 | 523 | 3 | 93 | Fibronectin type 3 domain; One of three types of internal repeats found in the plasma protein fibronectin. Its tenth fibronectin type III repeat contains an RGD cell recognition sequence in a flexible loop between 2 strands. Approximately 2% of all animal proteins contain the FN3 repeat; including extracellular and intracellular proteins, membrane spanning cytokine receptors, growth hormone receptors, tyrosine phosphatase receptors, and adhesion molecules. FN3-like domains are also found in bacterial glycosyl hydrolases. |
| PRK15319 | PRK15319 | 2.45e-08 | 549 | 991 | 583 | 1005 | fibronectin-binding autotransporter adhesin ShdA. |
| pfam18884 | TSP3_bac | 1.22e-06 | 383 | 403 | 1 | 21 | Bacterial TSP3 repeat. This entry contains a novel bacterial thrombospondin type 3 repeat which differs from the typical consensus by containing a glutamate in place of one of the calcium binding aspartate residues. |
| PRK15319 | PRK15319 | 6.16e-06 | 594 | 1071 | 802 | 1365 | fibronectin-binding autotransporter adhesin ShdA. |
| pfam00041 | fn3 | 6.75e-06 | 435 | 506 | 2 | 76 | Fibronectin type III domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AEV97716.1 | 2.12e-269 | 7 | 1099 | 11 | 1105 |
| QCD38445.1 | 1.36e-133 | 13 | 523 | 16 | 561 |
| QUT73817.1 | 2.77e-133 | 10 | 548 | 7 | 593 |
| QCP72135.1 | 1.48e-128 | 13 | 523 | 16 | 561 |
| CEI60018.1 | 7.12e-119 | 10 | 424 | 4 | 413 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4U3H_A | 2.52e-07 | 436 | 520 | 17 | 99 | Crystalstructure of FN3con [synthetic construct] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B0XMA2 | 1.56e-110 | 8 | 424 | 3 | 414 | Probable pectate lyase C OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=plyC PE=3 SV=1 |
| Q4WL88 | 2.18e-110 | 12 | 424 | 7 | 414 | Probable pectate lyase C OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=plyC PE=3 SV=1 |
| A1DPF0 | 8.22e-110 | 8 | 424 | 3 | 414 | Probable pectate lyase C OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=plyC PE=3 SV=1 |
| B8NQQ7 | 5.92e-108 | 25 | 424 | 21 | 413 | Probable pectate lyase C OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=plyC PE=3 SV=1 |
| Q2UB83 | 1.16e-106 | 25 | 424 | 21 | 413 | Probable pectate lyase C OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=plyC PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
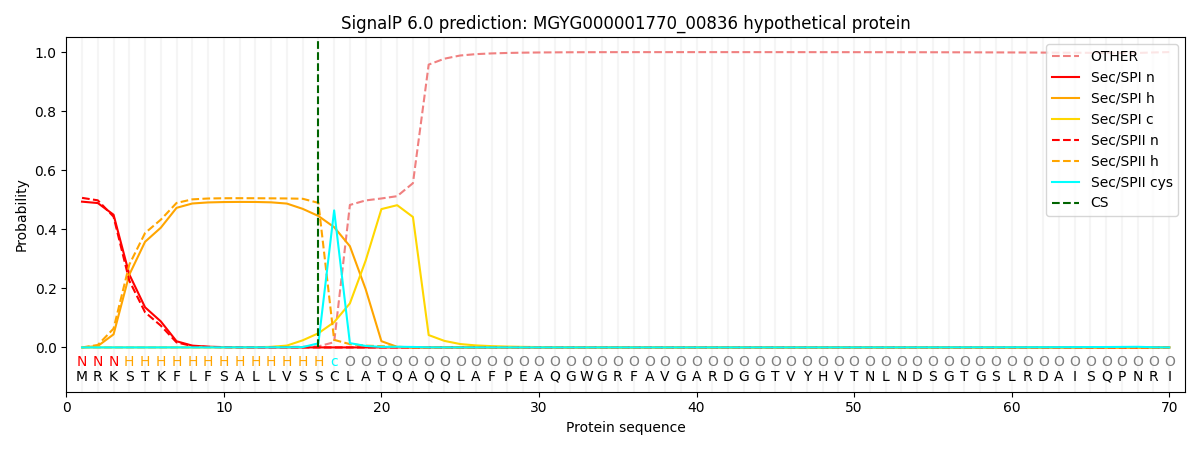
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000560 | 0.483477 | 0.515171 | 0.000281 | 0.000270 | 0.000219 |
