You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001770_01925
You are here: Home > Sequence: MGYG000001770_01925
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp900313215 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900313215 | |||||||||||
| CAZyme ID | MGYG000001770_01925 | |||||||||||
| CAZy Family | PL10 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 7517; End: 9619 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL10 | 50 | 339 | 4.8e-113 | 0.9826388888888888 |
| CE8 | 397 | 620 | 1.9e-27 | 0.7673611111111112 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam09492 | Pec_lyase | 6.21e-148 | 50 | 339 | 1 | 284 | Pectic acid lyase. Members of this family are isozymes of pectate lyase (EC:4.2.2.2), also called polygalacturonic transeliminase and alpha-1,4-D-endopolygalacturonic acid lyase. |
| TIGR02474 | pec_lyase | 8.92e-113 | 50 | 336 | 1 | 282 | pectate lyase, PelA/Pel-15E family. Members of this family are isozymes of pectate lyase (EC 4.2.2.2), also called polygalacturonic transeliminase and alpha-1,4-D-endopolygalacturonic acid lyase. [Energy metabolism, Biosynthesis and degradation of polysaccharides] |
| pfam01095 | Pectinesterase | 2.49e-07 | 398 | 445 | 15 | 64 | Pectinesterase. |
| PLN02432 | PLN02432 | 1.41e-06 | 397 | 491 | 25 | 109 | putative pectinesterase |
| PLN02176 | PLN02176 | 1.71e-06 | 398 | 550 | 54 | 193 | putative pectinesterase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ALJ61703.1 | 0.0 | 9 | 697 | 20 | 707 |
| QUT92735.1 | 0.0 | 9 | 700 | 20 | 710 |
| ALK83448.1 | 0.0 | 27 | 697 | 44 | 713 |
| QEW34712.1 | 0.0 | 27 | 697 | 44 | 713 |
| QJR78975.1 | 0.0 | 27 | 697 | 44 | 713 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1R76_A | 2.36e-60 | 33 | 350 | 77 | 404 | ChainA, pectate lyase [Niveispirillum irakense] |
| 1GXM_A | 1.32e-49 | 44 | 335 | 38 | 314 | Family10 polysaccharide lyase from Cellvibrio cellulosa [Cellvibrio japonicus],1GXM_B Family 10 polysaccharide lyase from Cellvibrio cellulosa [Cellvibrio japonicus],1GXN_A Family 10 polysaccharide lyase from Cellvibrio cellulosa [Cellvibrio japonicus] |
| 1GXO_A | 1.74e-48 | 44 | 335 | 38 | 314 | MutantD189A of Family 10 polysaccharide lyase from Cellvibrio cellulosa in complex with trigalaturonic acid [Cellvibrio japonicus] |
| 2NSP_A | 1.39e-08 | 398 | 599 | 21 | 205 | ChainA, Pectinesterase A [Dickeya dadantii 3937],2NSP_B Chain B, Pectinesterase A [Dickeya dadantii 3937],2NST_A Chain A, Pectinesterase A [Dickeya dadantii 3937],2NST_B Chain B, Pectinesterase A [Dickeya dadantii 3937],2NT6_A Chain A, Pectinesterase A [Dickeya dadantii 3937],2NT6_B Chain B, Pectinesterase A [Dickeya dadantii 3937],2NT9_A Chain A, Pectinesterase A [Dickeya dadantii 3937],2NT9_B Chain B, Pectinesterase A [Dickeya dadantii 3937] |
| 1QJV_A | 1.36e-07 | 398 | 489 | 21 | 109 | ChainA, PECTIN METHYLESTERASE [Dickeya chrysanthemi],1QJV_B Chain B, PECTIN METHYLESTERASE [Dickeya chrysanthemi] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P0C1A8 | 2.65e-07 | 398 | 489 | 45 | 133 | Pectinesterase A OS=Dickeya chrysanthemi OX=556 GN=pemA PE=1 SV=1 |
| P0C1A9 | 8.22e-07 | 398 | 489 | 45 | 133 | Pectinesterase A OS=Dickeya dadantii (strain 3937) OX=198628 GN=pemA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
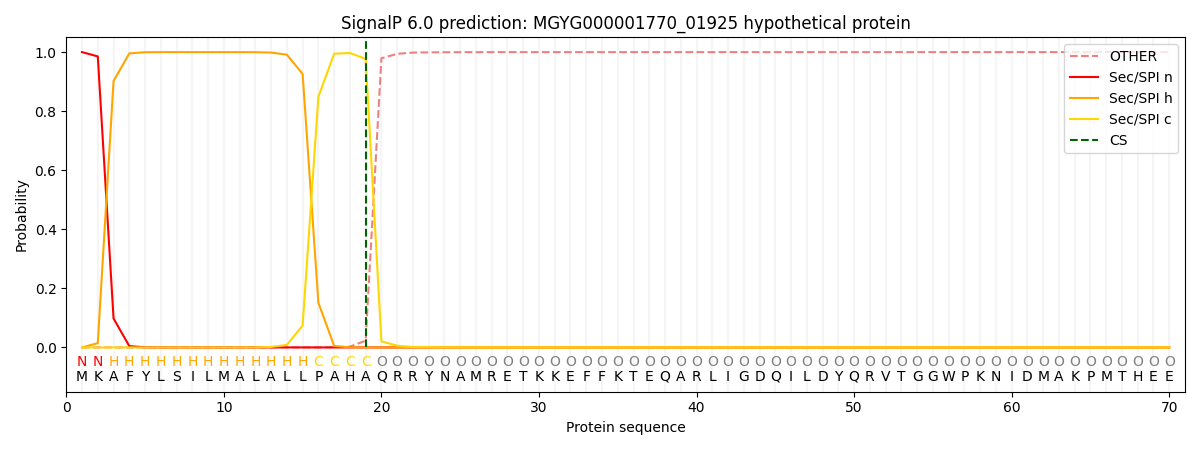
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000323 | 0.999048 | 0.000179 | 0.000145 | 0.000144 | 0.000130 |
