You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001779_02256
You are here: Home > Sequence: MGYG000001779_02256
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
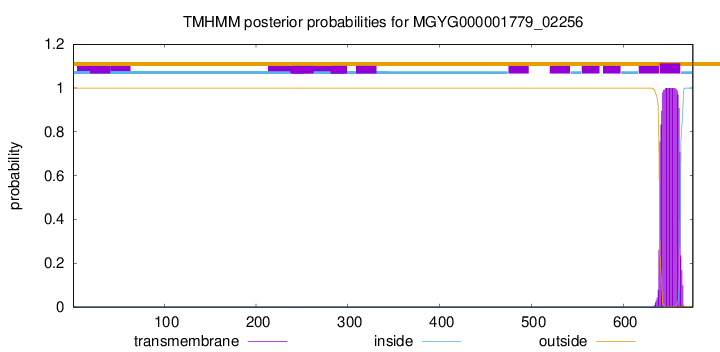
TMHMM annotations
Basic Information help
| Species | UMGS1851 sp900555605 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; UBA1212; UBA1255; UMGS1851; UMGS1851 sp900555605 | |||||||||||
| CAZyme ID | MGYG000001779_02256 | |||||||||||
| CAZy Family | GH32 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1780; End: 3810 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH32 | 115 | 442 | 6.7e-72 | 0.9795221843003413 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00640 | Glyco_32 | 4.32e-110 | 115 | 569 | 1 | 437 | Glycosyl hydrolases family 32. |
| cd18622 | GH32_Inu-like | 1.96e-99 | 123 | 434 | 4 | 289 | glycoside hydrolase family 32 protein such as Aspergillus ficuum endo-inulinase (Inu2). This subfamily of glycosyl hydrolase family GH32 includes endo-inulinase (inu2, EC 3.2.1.7), exo-inulinase (Inu1, EC 3.2.1.80), invertase (EC 3.2.1.26), and levan fructotransferase (LftA, EC 4.2.2.16), among others. These enzymes cleave sucrose into fructose and glucose via beta-fructofuranosidase activity, producing invert sugar that is a mixture of dextrorotatory D-glucose and levorotatory D-fructose, thus named invertase (EC 3.2.1.26). These retaining enzymes (i.e. they retain the configuration at anomeric carbon atom of the substrate) catalyze hydrolysis in two steps involving a covalent glycosyl enzyme intermediate: an aspartate located close to the N-terminus acts as the catalytic nucleophile and a glutamate acts as the general acid/base; a conserved aspartate residue in the Arg-Asp-Pro (RDP) motif stabilizes the transition state. These enzymes are predicted to display a 5-fold beta-propeller fold as found for GH43 and CH68. The breakdown of sucrose is widely used as a carbon or energy source by bacteria, fungi, and plants. Invertase is used commercially in the confectionery industry, since fructose has a sweeter taste than sucrose and a lower tendency to crystallize. A common structural feature of all these enzymes is a 5-bladed beta-propeller domain, similar to GH43, that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| COG1621 | SacC | 1.04e-97 | 106 | 607 | 24 | 486 | Sucrose-6-phosphate hydrolase SacC, GH32 family [Carbohydrate transport and metabolism]. |
| pfam00251 | Glyco_hydro_32N | 5.24e-81 | 115 | 446 | 1 | 307 | Glycosyl hydrolases family 32 N-terminal domain. This domain corresponds to the N-terminal domain of glycosyl hydrolase family 32 which forms a five bladed beta propeller structure. |
| cd08996 | GH32_FFase | 1.88e-73 | 123 | 434 | 3 | 281 | Glycosyl hydrolase family 32, beta-fructosidases. Glycosyl hydrolase family GH32 cleaves sucrose into fructose and glucose via beta-fructofuranosidase activity, producing invert sugar that is a mixture of dextrorotatory D-glucose and levorotatory D-fructose, thus named invertase (EC 3.2.1.26). This family also contains other fructofuranosidases such as inulinase (EC 3.2.1.7), exo-inulinase (EC 3.2.1.80), levanase (EC 3.2.1.65), and transfructosidases such sucrose:sucrose 1-fructosyltransferase (EC 2.4.1.99), fructan:fructan 1-fructosyltransferase (EC 2.4.1.100), sucrose:fructan 6-fructosyltransferase (EC 2.4.1.10), fructan:fructan 6G-fructosyltransferase (EC 2.4.1.243) and levan fructosyltransferases (EC 2.4.1.-). These retaining enzymes (i.e. they retain the configuration at anomeric carbon atom of the substrate) catalyze hydrolysis in two steps involving a covalent glycosyl enzyme intermediate: an aspartate located close to the N-terminus acts as the catalytic nucleophile and a glutamate acts as the general acid/base; a conserved aspartate residue in the Arg-Asp-Pro (RDP) motif stabilizes the transition state. These enzymes are predicted to display a 5-fold beta-propeller fold as found for GH43 and CH68. The breakdown of sucrose is widely used as a carbon or energy source by bacteria, fungi, and plants. Invertase is used commercially in the confectionery industry, since fructose has a sweeter taste than sucrose and a lower tendency to crystallize. A common structural feature of all these enzymes is a 5-bladed beta-propeller domain, similar to GH43, that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QJA04153.1 | 1.08e-117 | 20 | 609 | 423 | 1000 |
| QIX10455.1 | 6.07e-114 | 21 | 609 | 425 | 1000 |
| BCT45591.1 | 3.85e-113 | 87 | 609 | 483 | 984 |
| AGK95925.1 | 4.67e-113 | 2 | 626 | 381 | 992 |
| AJA50828.1 | 7.42e-107 | 2 | 611 | 379 | 970 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3RWK_X | 7.12e-57 | 102 | 602 | 20 | 510 | Firstcrystal structure of an endo-inulinase, from Aspergillus ficuum: structural analysis and comparison with other GH32 enzymes. [Aspergillus ficuum],3SC7_X First crystal structure of an endo-inulinase, from Aspergillus ficuum: structural analysis and comparison with other GH32 enzymes. [Aspergillus ficuum] |
| 1Y4W_A | 4.50e-53 | 106 | 609 | 3 | 517 | Crystalstructure of exo-inulinase from Aspergillus awamori in spacegroup P21 [Aspergillus awamori],1Y9G_A Crystal structure of exo-inulinase from Aspergillus awamori complexed with fructose [Aspergillus awamori],1Y9M_A Crystal structure of exo-inulinase from Aspergillus awamori in spacegroup P212121 [Aspergillus awamori] |
| 3KF3_A | 4.25e-46 | 108 | 408 | 7 | 288 | ChainA, Invertase [Schwanniomyces occidentalis],3KF3_B Chain B, Invertase [Schwanniomyces occidentalis] |
| 3KF5_A | 4.48e-46 | 108 | 408 | 10 | 291 | ChainA, Invertase [Schwanniomyces occidentalis],3KF5_B Chain B, Invertase [Schwanniomyces occidentalis] |
| 3U75_A | 4.34e-45 | 108 | 408 | 33 | 314 | ChainA, Fructofuranosidase [Schwanniomyces occidentalis],3U75_B Chain B, Fructofuranosidase [Schwanniomyces occidentalis],3U75_C Chain C, Fructofuranosidase [Schwanniomyces occidentalis],3U75_D Chain D, Fructofuranosidase [Schwanniomyces occidentalis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P05656 | 1.41e-95 | 106 | 609 | 30 | 513 | Levanase OS=Bacillus subtilis (strain 168) OX=224308 GN=sacC PE=1 SV=1 |
| O31411 | 6.37e-92 | 103 | 609 | 390 | 879 | Levanase (Fragment) OS=Bacillus sp. (strain L7) OX=62626 PE=1 SV=2 |
| Q03174 | 6.69e-71 | 49 | 617 | 380 | 923 | Fructan beta-fructosidase OS=Streptococcus mutans serotype c (strain ATCC 700610 / UA159) OX=210007 GN=fruA PE=2 SV=2 |
| A5ABL2 | 1.52e-57 | 102 | 602 | 20 | 510 | Extracellular endo-inulinase inuA OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=inuA PE=1 SV=1 |
| O74641 | 7.70e-57 | 102 | 602 | 20 | 510 | Extracellular endo-inulinase inuA OS=Aspergillus niger OX=5061 GN=inuA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
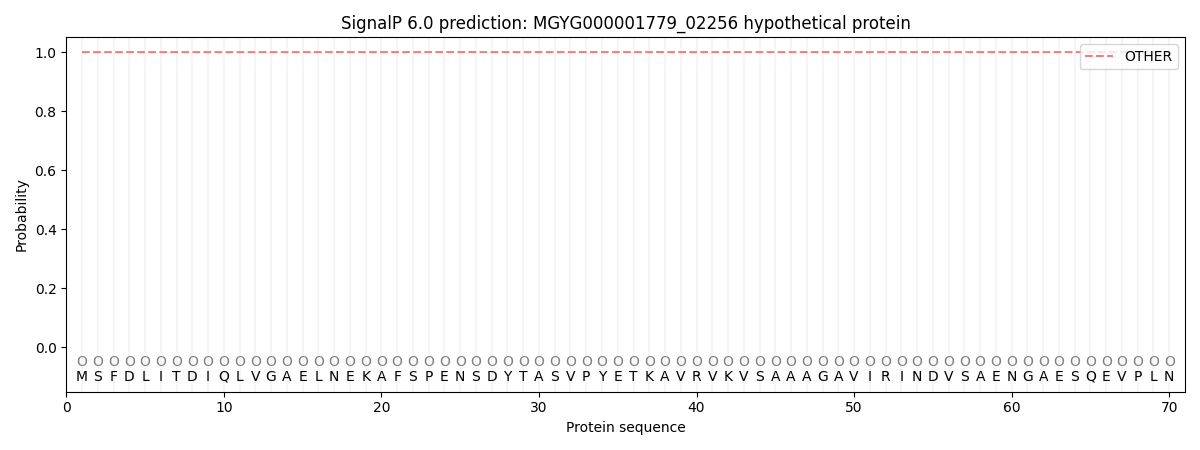
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000011 | 0.000029 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |