You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001780_01237
You are here: Home > Sequence: MGYG000001780_01237
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; | |||||||||||
| CAZyme ID | MGYG000001780_01237 | |||||||||||
| CAZy Family | GH105 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 40379; End: 42484 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH105 | 51 | 395 | 6.2e-123 | 0.9819277108433735 |
| CE8 | 410 | 690 | 1.4e-101 | 0.9861111111111112 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam07470 | Glyco_hydro_88 | 4.89e-135 | 32 | 396 | 1 | 342 | Glycosyl Hydrolase Family 88. Unsaturated glucuronyl hydrolase catalyzes the hydrolytic release of unsaturated glucuronic acids from oligosaccharides (EC:3.2.1.-) produced by the reactions of polysaccharide lyases. |
| COG4225 | YesR | 1.45e-107 | 52 | 396 | 28 | 356 | Rhamnogalacturonyl hydrolase YesR [Carbohydrate transport and metabolism]. |
| PLN02773 | PLN02773 | 2.57e-76 | 412 | 690 | 9 | 296 | pectinesterase |
| pfam01095 | Pectinesterase | 4.75e-75 | 410 | 690 | 2 | 295 | Pectinesterase. |
| PLN02682 | PLN02682 | 5.51e-68 | 409 | 690 | 70 | 361 | pectinesterase family protein |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ADY37138.1 | 0.0 | 8 | 699 | 4 | 698 |
| QDO69177.1 | 3.05e-293 | 1 | 396 | 1 | 396 |
| QUT93209.1 | 6.14e-293 | 1 | 396 | 1 | 396 |
| ALJ61262.1 | 6.14e-293 | 1 | 396 | 1 | 396 |
| QUT77882.1 | 1.58e-272 | 1 | 396 | 1 | 396 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4WU0_A | 1.18e-117 | 58 | 396 | 23 | 361 | StructuralAnalysis of C. acetobutylicum ATCC 824 Glycoside Hydrolase From Family 105 [Clostridium acetobutylicum ATCC 824],4WU0_B Structural Analysis of C. acetobutylicum ATCC 824 Glycoside Hydrolase From Family 105 [Clostridium acetobutylicum ATCC 824] |
| 1NC5_A | 6.85e-81 | 51 | 396 | 32 | 367 | Structureof Protein of Unknown Function of YteR from Bacillus Subtilis [Bacillus subtilis],2D8L_A Crystal Structure of Unsaturated Rhamnogalacturonyl Hydrolase in complex with dGlcA-GalNAc [Bacillus subtilis] |
| 2GH4_A | 2.74e-80 | 51 | 396 | 22 | 357 | ChainA, Putative glycosyl hydrolase yteR [Bacillus subtilis] |
| 1GQ8_A | 3.07e-36 | 410 | 683 | 9 | 294 | Pectinmethylesterase from Carrot [Daucus carota] |
| 1XG2_A | 5.71e-34 | 411 | 672 | 6 | 277 | ChainA, Pectinesterase 1 [Solanum lycopersicum] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O34559 | 3.75e-80 | 51 | 396 | 32 | 367 | Unsaturated rhamnogalacturonyl hydrolase YteR OS=Bacillus subtilis (strain 168) OX=224308 GN=yteR PE=1 SV=1 |
| Q9LVQ0 | 9.38e-49 | 412 | 690 | 9 | 296 | Pectinesterase 31 OS=Arabidopsis thaliana OX=3702 GN=PME31 PE=1 SV=1 |
| Q9FJ21 | 1.77e-45 | 339 | 690 | 196 | 553 | Probable pectinesterase/pectinesterase inhibitor 58 OS=Arabidopsis thaliana OX=3702 GN=PME58 PE=2 SV=1 |
| Q8GXA1 | 1.09e-44 | 397 | 685 | 251 | 546 | Probable pectinesterase/pectinesterase inhibitor 23 OS=Arabidopsis thaliana OX=3702 GN=PME23 PE=2 SV=3 |
| P41510 | 1.20e-43 | 407 | 683 | 270 | 557 | Probable pectinesterase/pectinesterase inhibitor OS=Brassica napus OX=3708 GN=BP19 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
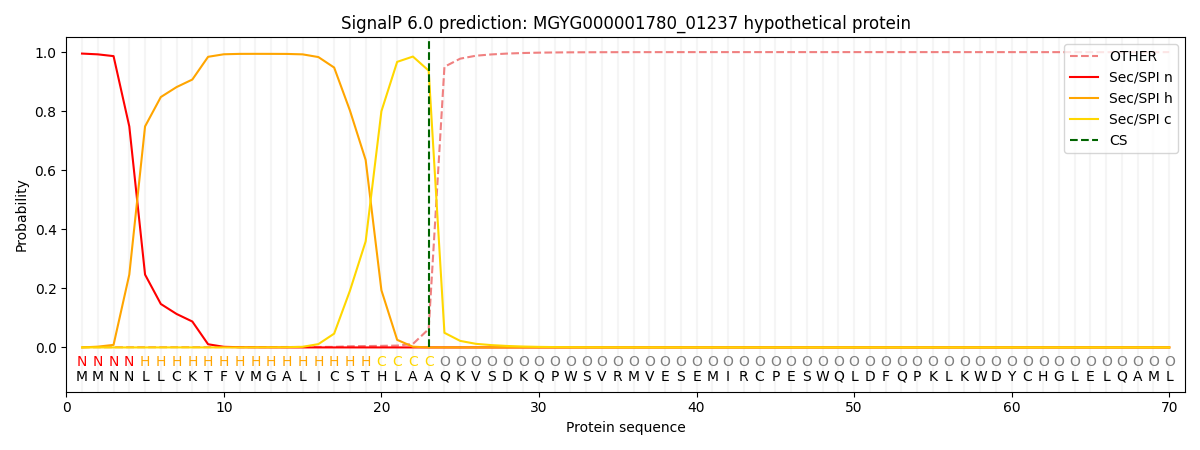
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001922 | 0.992189 | 0.005172 | 0.000285 | 0.000224 | 0.000197 |
