You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001789_01600
You are here: Home > Sequence: MGYG000001789_01600
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phocaeicola sp002161565 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Phocaeicola; Phocaeicola sp002161565 | |||||||||||
| CAZyme ID | MGYG000001789_01600 | |||||||||||
| CAZy Family | GH28 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 31882; End: 33297 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH28 | 55 | 405 | 2.3e-68 | 0.9323076923076923 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG5434 | Pgu1 | 1.21e-59 | 30 | 311 | 82 | 390 | Polygalacturonase [Carbohydrate transport and metabolism]. |
| PLN03010 | PLN03010 | 2.46e-24 | 31 | 378 | 47 | 379 | polygalacturonase |
| PLN02188 | PLN02188 | 5.67e-23 | 28 | 375 | 34 | 348 | polygalacturonase/glycoside hydrolase family protein |
| PLN02218 | PLN02218 | 2.46e-20 | 32 | 317 | 69 | 343 | polygalacturonase ADPG |
| PLN02793 | PLN02793 | 3.05e-17 | 30 | 317 | 52 | 328 | Probable polygalacturonase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ADY36938.1 | 0.0 | 1 | 471 | 1 | 471 |
| ALJ60362.1 | 4.12e-253 | 12 | 469 | 7 | 467 |
| QUT88631.1 | 1.18e-252 | 12 | 469 | 7 | 467 |
| QDO68557.1 | 1.67e-252 | 6 | 469 | 2 | 467 |
| QEW34965.1 | 3.31e-243 | 10 | 469 | 13 | 467 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5OLP_A | 3.27e-25 | 31 | 292 | 45 | 330 | Galacturonidase[Bacteroides thetaiotaomicron VPI-5482],5OLP_B Galacturonidase [Bacteroides thetaiotaomicron VPI-5482] |
| 3JUR_A | 6.80e-19 | 32 | 311 | 29 | 343 | Thecrystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_B The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_C The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_D The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima] |
| 4C2L_A | 3.85e-16 | 38 | 317 | 23 | 292 | Crystalstructure of endo-xylogalacturonan hydrolase from Aspergillus tubingensis [Aspergillus tubingensis] |
| 2UVE_A | 1.78e-13 | 25 | 238 | 151 | 405 | Structureof Yersinia enterocolitica Family 28 Exopolygalacturonase [Yersinia enterocolitica],2UVE_B Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase [Yersinia enterocolitica],2UVF_A Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase in Complex with Digalaturonic Acid [Yersinia enterocolitica],2UVF_B Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase in Complex with Digalaturonic Acid [Yersinia enterocolitica] |
| 2PYG_A | 6.44e-08 | 31 | 70 | 3 | 42 | Azotobactervinelandii Mannuronan C-5 epimerase AlgE4 A-module [Azotobacter vinelandii],2PYG_B Azotobacter vinelandii Mannuronan C-5 epimerase AlgE4 A-module [Azotobacter vinelandii],2PYH_A Azotobacter vinelandii Mannuronan C-5 epimerase AlgE4 A-module complexed with mannuronan trisaccharide [Azotobacter vinelandii],2PYH_B Azotobacter vinelandii Mannuronan C-5 epimerase AlgE4 A-module complexed with mannuronan trisaccharide [Azotobacter vinelandii] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A7PZL3 | 2.22e-25 | 22 | 299 | 57 | 339 | Probable polygalacturonase OS=Vitis vinifera OX=29760 GN=GSVIVT00026920001 PE=1 SV=1 |
| Q8RY29 | 6.84e-21 | 22 | 317 | 61 | 343 | Polygalacturonase ADPG2 OS=Arabidopsis thaliana OX=3702 GN=ADPG2 PE=2 SV=2 |
| Q9LW07 | 9.54e-17 | 27 | 317 | 20 | 289 | Probable polygalacturonase At3g15720 OS=Arabidopsis thaliana OX=3702 GN=At3g15720 PE=3 SV=1 |
| P35339 | 9.83e-17 | 31 | 314 | 41 | 350 | Exopolygalacturonase OS=Zea mays OX=4577 GN=PG2C PE=2 SV=1 |
| Q7M1E7 | 1.19e-16 | 30 | 314 | 58 | 331 | Polygalacturonase OS=Chamaecyparis obtusa OX=13415 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
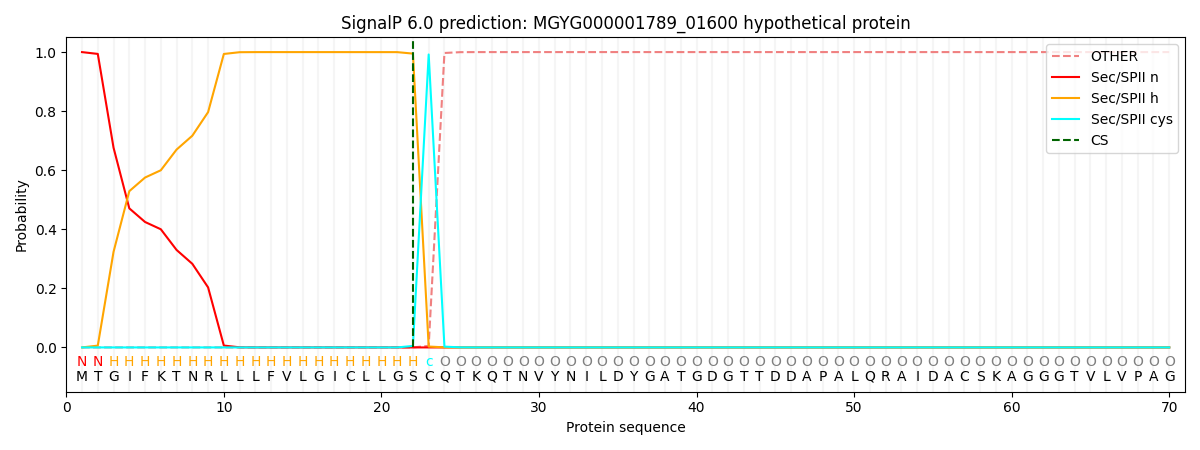
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000011 | 1.000018 | 0.000000 | 0.000000 | 0.000000 |
