You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001803_01545
You are here: Home > Sequence: MGYG000001803_01545
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
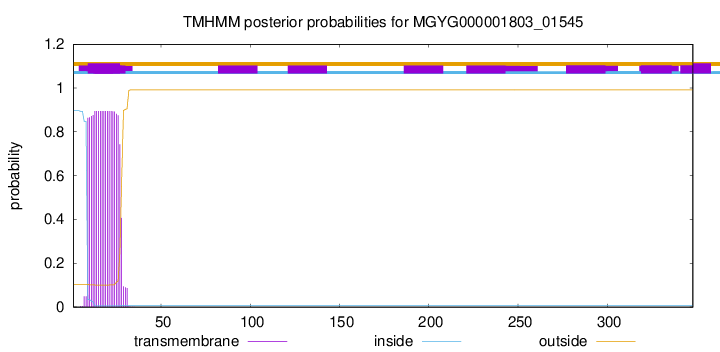
TMHMM annotations
Basic Information help
| Species | RC9 sp900546925 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; UBA932; RC9; RC9 sp900546925 | |||||||||||
| CAZyme ID | MGYG000001803_01545 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | Membrane-bound lytic murein transglycosylase F | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 12618; End: 13664 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH23 | 192 | 341 | 4.3e-25 | 0.7925925925925926 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd13403 | MLTF-like | 1.33e-50 | 184 | 343 | 3 | 159 | membrane-bound lytic murein transglycosylase F (MLTF) and similar proteins. This subfamily includes membrane-bound lytic murein transglycosylase F (MltF, murein lyase F) that degrades murein glycan strands. It is responsible for catalyzing the release of 1,6-anhydromuropeptides from peptidoglycan. Lytic transglycosylase catalyzes the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc) as do goose-type lysozymes. However, in addition, it also makes a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
| cd00254 | LT-like | 1.45e-28 | 193 | 283 | 1 | 87 | lytic transglycosylase(LT)-like domain. Members include the soluble and insoluble membrane-bound LTs in bacteria and LTs in bacteriophage lambda. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
| cd16896 | LT_Slt70-like | 8.89e-28 | 177 | 282 | 3 | 110 | uncharacterized lytic transglycosylase subfamily with similarity to Slt70. Uncharacterized lytic transglycosylase (LT) with a conserved sequence pattern suggesting similarity to the Slt70, a 70kda soluble lytic transglycosylase which also has an N-terminal U-shaped U-domain and a linker L-domain. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
| PRK10859 | PRK10859 | 1.94e-25 | 178 | 335 | 290 | 442 | membrane-bound lytic murein transglycosylase MltF. |
| COG4623 | MltF | 5.16e-25 | 178 | 338 | 271 | 426 | Membrane-bound lytic murein transglycosylase MltF [Cell wall/membrane/envelope biogenesis, Signal transduction mechanisms]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CRY96252.1 | 2.52e-122 | 1 | 293 | 1 | 294 |
| AGY53417.1 | 2.54e-45 | 38 | 337 | 37 | 341 |
| QZT36545.1 | 5.70e-33 | 34 | 346 | 149 | 474 |
| QZE14494.1 | 3.93e-32 | 76 | 346 | 195 | 474 |
| QKG79130.1 | 6.38e-32 | 83 | 346 | 192 | 462 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7EYB_I | 3.48e-08 | 173 | 281 | 6 | 115 | ChainI, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7EYB_J Chain J, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7EYB_K Chain K, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7EYB_L Chain L, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7K5C_B Chain B, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7K5C_D Chain D, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7K5C_F Chain F, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7K5C_G Chain G, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7K5C_J Chain J, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7K5C_L Chain L, Peptidoglycan transglycosylase gp16 [Escherichia phage T7] |
| 6YT5_G | 3.49e-08 | 173 | 281 | 28 | 137 | ChainG, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],6YT5_H Chain H, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],6YT5_I Chain I, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],6YT5_J Chain J, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],6YT5_K Chain K, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],6YT5_L Chain L, Peptidoglycan transglycosylase gp16 [Escherichia phage T7] |
| 1QSA_A | 1.55e-07 | 178 | 289 | 453 | 567 | CrystalStructure Of The 70 Kda Soluble Lytic Transglycosylase Slt70 From Escherichia Coli At 1.65 Angstroms Resolution [Escherichia coli],1QTE_A Crystal Structure Of The 70 Kda Soluble Lytic Transglycosylase Slt70 From Escherichia Coli At 1.90 A Resolution In Complex With A 1,6- Anhydromurotripeptide [Escherichia coli] |
| 1SLY_A | 2.73e-07 | 178 | 289 | 453 | 567 | ComplexOf The 70-Kda Soluble Lytic Transglycosylase With Bulgecin A [Escherichia coli] |
| 6FBT_A | 3.60e-07 | 180 | 282 | 451 | 555 | ChainA, Lytic murein transglycosylase [Pseudomonas aeruginosa] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A8ZWR8 | 1.52e-24 | 83 | 341 | 182 | 446 | Membrane-bound lytic murein transglycosylase F OS=Desulfococcus oleovorans (strain DSM 6200 / JCM 39069 / Hxd3) OX=96561 GN=mltF PE=3 SV=2 |
| Q0I361 | 8.27e-20 | 46 | 338 | 151 | 443 | Membrane-bound lytic murein transglycosylase F OS=Haemophilus somnus (strain 129Pt) OX=205914 GN=mltF PE=3 SV=2 |
| B0UUK3 | 2.02e-19 | 46 | 338 | 151 | 443 | Membrane-bound lytic murein transglycosylase F OS=Histophilus somni (strain 2336) OX=228400 GN=mltF PE=3 SV=1 |
| B4RVK5 | 4.65e-19 | 175 | 342 | 291 | 453 | Membrane-bound lytic murein transglycosylase F OS=Alteromonas mediterranea (strain DSM 17117 / CIP 110805 / LMG 28347 / Deep ecotype) OX=1774373 GN=mltF PE=3 SV=3 |
| Q4QNV4 | 6.80e-19 | 63 | 345 | 165 | 451 | Membrane-bound lytic murein transglycosylase F OS=Haemophilus influenzae (strain 86-028NP) OX=281310 GN=mltF PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
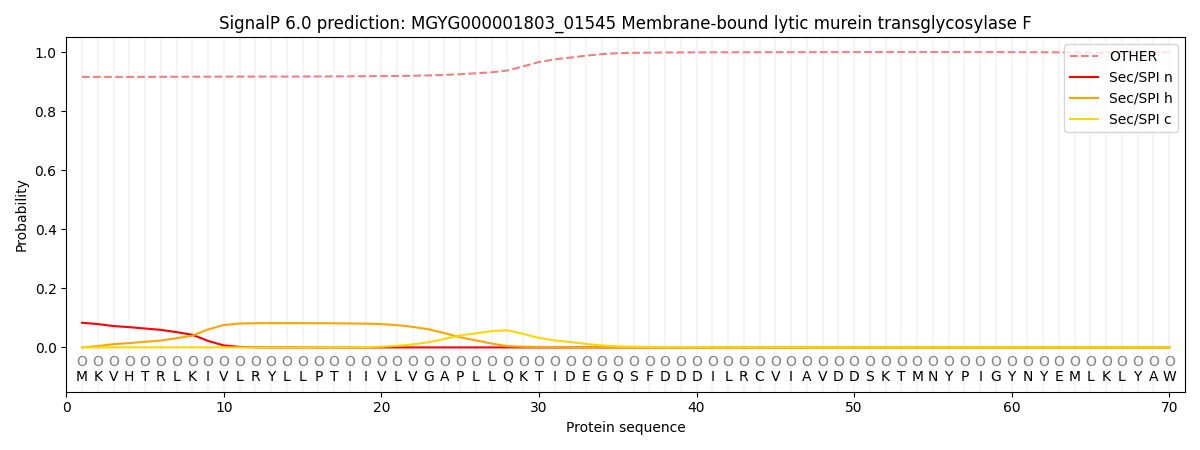
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.920010 | 0.078655 | 0.000492 | 0.000203 | 0.000152 | 0.000509 |