You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001814_01038
You are here: Home > Sequence: MGYG000001814_01038
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | COE1 sp900753305 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; COE1; COE1 sp900753305 | |||||||||||
| CAZyme ID | MGYG000001814_01038 | |||||||||||
| CAZy Family | CE2 | |||||||||||
| CAZyme Description | Cellulase/esterase CelE | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 65052; End: 66152 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE2 | 129 | 362 | 1.2e-61 | 0.9952153110047847 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd01831 | Endoglucanase_E_like | 3.08e-33 | 129 | 363 | 1 | 169 | Endoglucanase E-like members of the SGNH hydrolase family; Endoglucanase E catalyzes the endohydrolysis of 1,4-beta-glucosidic linkages in cellulose, lichenin and cereal beta-D-glucans. |
| cd00229 | SGNH_hydrolase | 4.44e-06 | 132 | 298 | 3 | 111 | SGNH_hydrolase, or GDSL_hydrolase, is a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the typical Ser-His-Asp(Glu) triad from other serine hydrolases, but may lack the carboxlic acid. |
| cd01827 | sialate_O-acetylesterase_like1 | 6.05e-04 | 227 | 300 | 66 | 115 | sialate O-acetylesterase_like family of the SGNH hydrolases, a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the Ser-His-Asp(Glu) triad found in other serine hydrolases. |
| cd01834 | SGNH_hydrolase_like_2 | 0.002 | 129 | 300 | 3 | 111 | SGNH_hydrolase subfamily. SGNH hydrolases are a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the Ser-His-Asp(Glu) triad found in other serine hydrolases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| APO43020.1 | 1.40e-158 | 1 | 366 | 8 | 373 |
| QZN77000.1 | 8.06e-158 | 4 | 366 | 11 | 373 |
| AYQ74824.1 | 1.32e-156 | 4 | 366 | 11 | 373 |
| QKS57722.1 | 6.64e-155 | 4 | 366 | 13 | 375 |
| ASA24859.1 | 6.68e-154 | 4 | 366 | 9 | 371 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3U37_A | 3.61e-116 | 3 | 366 | 39 | 404 | AnAcetyl Xylan Esterase (Est2A) from the Rumen Bacterium Butyrivibrio proteoclasticus. [Butyrivibrio proteoclasticus B316],3U37_B An Acetyl Xylan Esterase (Est2A) from the Rumen Bacterium Butyrivibrio proteoclasticus. [Butyrivibrio proteoclasticus B316],3U37_C An Acetyl Xylan Esterase (Est2A) from the Rumen Bacterium Butyrivibrio proteoclasticus. [Butyrivibrio proteoclasticus B316],3U37_D An Acetyl Xylan Esterase (Est2A) from the Rumen Bacterium Butyrivibrio proteoclasticus. [Butyrivibrio proteoclasticus B316],3U37_E An Acetyl Xylan Esterase (Est2A) from the Rumen Bacterium Butyrivibrio proteoclasticus. [Butyrivibrio proteoclasticus B316],3U37_F An Acetyl Xylan Esterase (Est2A) from the Rumen Bacterium Butyrivibrio proteoclasticus. [Butyrivibrio proteoclasticus B316],3U37_G An Acetyl Xylan Esterase (Est2A) from the Rumen Bacterium Butyrivibrio proteoclasticus. [Butyrivibrio proteoclasticus B316],3U37_H An Acetyl Xylan Esterase (Est2A) from the Rumen Bacterium Butyrivibrio proteoclasticus. [Butyrivibrio proteoclasticus B316] |
| 4DEV_A | 1.17e-114 | 3 | 366 | 39 | 404 | AnAcetyl Xylan Esterase (Est2A) from the Rumen Bacterium Butyrivibrio proteoclasticus. [Butyrivibrio proteoclasticus B316],4DEV_B An Acetyl Xylan Esterase (Est2A) from the Rumen Bacterium Butyrivibrio proteoclasticus. [Butyrivibrio proteoclasticus B316],4DEV_C An Acetyl Xylan Esterase (Est2A) from the Rumen Bacterium Butyrivibrio proteoclasticus. [Butyrivibrio proteoclasticus B316],4DEV_D An Acetyl Xylan Esterase (Est2A) from the Rumen Bacterium Butyrivibrio proteoclasticus. [Butyrivibrio proteoclasticus B316],4DEV_E An Acetyl Xylan Esterase (Est2A) from the Rumen Bacterium Butyrivibrio proteoclasticus. [Butyrivibrio proteoclasticus B316],4DEV_F An Acetyl Xylan Esterase (Est2A) from the Rumen Bacterium Butyrivibrio proteoclasticus. [Butyrivibrio proteoclasticus B316],4DEV_G An Acetyl Xylan Esterase (Est2A) from the Rumen Bacterium Butyrivibrio proteoclasticus. [Butyrivibrio proteoclasticus B316],4DEV_H An Acetyl Xylan Esterase (Est2A) from the Rumen Bacterium Butyrivibrio proteoclasticus. [Butyrivibrio proteoclasticus B316] |
| 2WAO_A | 1.46e-17 | 28 | 366 | 29 | 331 | ChainA, ENDOGLUCANASE E [Acetivibrio thermocellus] |
| 2WAB_A | 3.64e-17 | 28 | 366 | 29 | 331 | ChainA, ENDOGLUCANASE E [Acetivibrio thermocellus] |
| 2W9X_A | 2.90e-08 | 129 | 242 | 144 | 250 | Theactive site of a carbohydrate esterase displays divergent catalytic and non-catalytic binding functions [Cellvibrio japonicus],2W9X_B The active site of a carbohydrate esterase displays divergent catalytic and non-catalytic binding functions [Cellvibrio japonicus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P10477 | 2.71e-16 | 28 | 366 | 510 | 812 | Cellulase/esterase CelE OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celE PE=1 SV=2 |
| B3PDE5 | 1.56e-07 | 129 | 242 | 144 | 250 | Acetylxylan esterase / glucomannan deacetylase OS=Cellvibrio japonicus (strain Ueda107) OX=498211 GN=ce2C PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
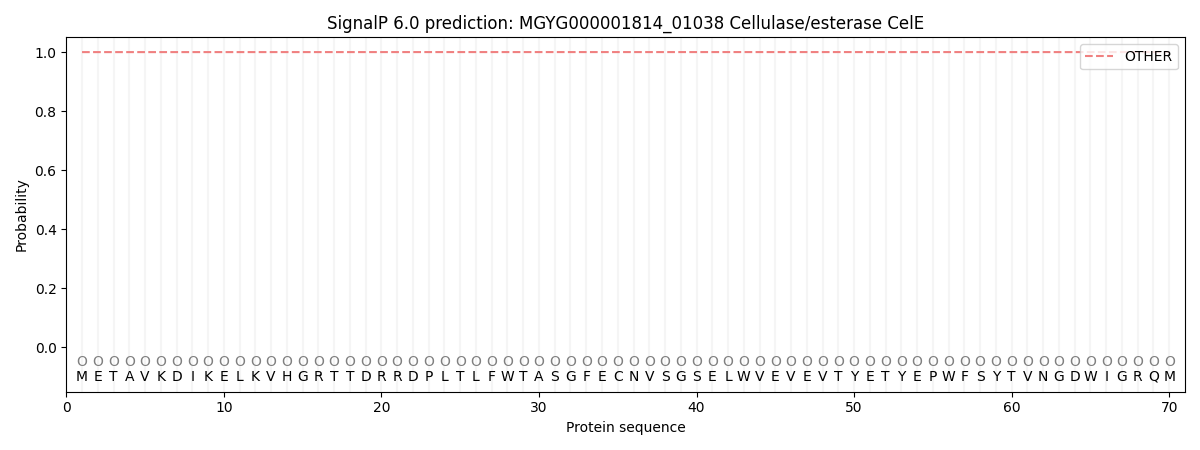
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000045 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
