You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001838_01050
You are here: Home > Sequence: MGYG000001838_01050
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Zag1 sp000438175 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Cyanobacteria; Vampirovibrionia; Gastranaerophilales; Gastranaerophilaceae; Zag1; Zag1 sp000438175 | |||||||||||
| CAZyme ID | MGYG000001838_01050 | |||||||||||
| CAZy Family | GH77 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 48635; End: 50638 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH77 | 62 | 598 | 1.8e-50 | 0.9230769230769231 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam02446 | Glyco_hydro_77 | 8.78e-25 | 97 | 366 | 46 | 282 | 4-alpha-glucanotransferase. These enzymes EC:2.4.1.25 transfer a segment of a (1,4)-alpha-D-glucan to a new 4-position in an acceptor, which may be glucose or (1,4)-alpha-D-glucan. |
| PRK14510 | PRK14510 | 5.88e-22 | 97 | 380 | 778 | 1046 | bifunctional glycogen debranching protein GlgX/4-alpha-glucanotransferase. |
| PRK14508 | PRK14508 | 1.60e-20 | 99 | 366 | 57 | 302 | 4-alpha-glucanotransferase; Provisional |
| COG1640 | MalQ | 1.93e-20 | 72 | 600 | 44 | 498 | 4-alpha-glucanotransferase [Carbohydrate transport and metabolism]. |
| TIGR00217 | malQ | 9.28e-16 | 99 | 600 | 67 | 497 | 4-alpha-glucanotransferase. This enzyme is known as amylomaltase and disproportionating enzyme. [Energy metabolism, Biosynthesis and degradation of polysaccharides] |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AOR39032.1 | 0.0 | 1 | 665 | 1 | 662 |
| AOR38234.1 | 2.07e-56 | 25 | 627 | 17 | 608 |
| QRK12034.1 | 1.14e-41 | 34 | 626 | 20 | 629 |
| QRN96421.1 | 2.95e-40 | 34 | 626 | 20 | 620 |
| AKT37259.1 | 3.39e-39 | 28 | 627 | 12 | 622 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q59266 | 1.11e-11 | 62 | 365 | 15 | 289 | 4-alpha-glucanotransferase OS=Clostridium butyricum OX=1492 GN=malQ PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
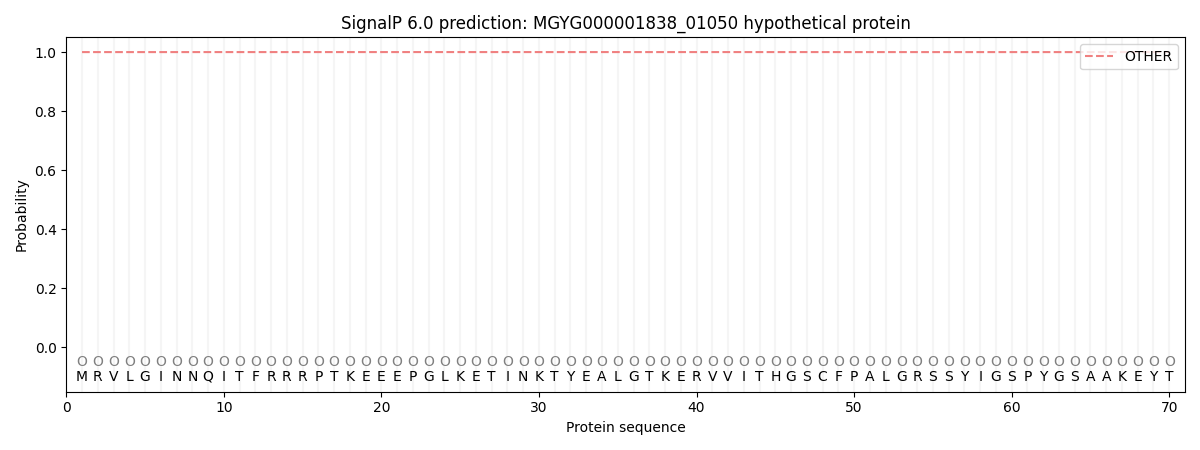
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000064 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
