You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001848_01856
You are here: Home > Sequence: MGYG000001848_01856
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Eubacterium_F sp000433735 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Eubacterium_F; Eubacterium_F sp000433735 | |||||||||||
| CAZyme ID | MGYG000001848_01856 | |||||||||||
| CAZy Family | GH73 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 10354; End: 12228 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG4193 | LytD | 2.02e-17 | 364 | 612 | 28 | 244 | Beta- N-acetylglucosaminidase [Carbohydrate transport and metabolism]. |
| pfam08239 | SH3_3 | 1.86e-10 | 40 | 95 | 1 | 54 | Bacterial SH3 domain. |
| PRK13914 | PRK13914 | 1.23e-08 | 37 | 129 | 84 | 194 | invasion associated endopeptidase. |
| smart00287 | SH3b | 4.35e-08 | 32 | 90 | 1 | 57 | Bacterial SH3 domain homologues. |
| COG3103 | YgiM | 1.80e-06 | 13 | 106 | 4 | 98 | Uncharacterized conserved protein YgiM, contains N-terminal SH3 domain, DUF1202 family [General function prediction only]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QHQ61172.1 | 4.14e-128 | 2 | 624 | 3 | 676 |
| BBF45313.1 | 2.75e-127 | 4 | 624 | 5 | 652 |
| BCN29841.1 | 2.64e-114 | 8 | 624 | 5 | 798 |
| ABX43817.1 | 3.72e-111 | 26 | 623 | 531 | 1182 |
| CUH91961.1 | 5.04e-111 | 37 | 623 | 144 | 816 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2KRS_A | 1.25e-07 | 35 | 96 | 3 | 61 | SolutionNMR structure of SH3 domain from CPF_0587 (fragment 415-479) from Clostridium perfringens. Northeast Structural Genomics Consortium (NESG) Target CpR74A. [Clostridium perfringens] |
| 6FXO_A | 1.38e-07 | 459 | 612 | 94 | 243 | ChainA, Bifunctional autolysin [Staphylococcus aureus subsp. aureus Mu50] |
| 2KQ8_A | 7.41e-07 | 37 | 95 | 7 | 61 | ChainA, Cell wall hydrolase [[Bacillus thuringiensis] serovar konkukian] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P39848 | 8.36e-11 | 385 | 612 | 679 | 879 | Beta-N-acetylglucosaminidase OS=Bacillus subtilis (strain 168) OX=224308 GN=lytD PE=1 SV=1 |
| Q8CPQ1 | 1.49e-09 | 385 | 612 | 1128 | 1334 | Bifunctional autolysin OS=Staphylococcus epidermidis (strain ATCC 12228 / FDA PCI 1200) OX=176280 GN=atl PE=3 SV=1 |
| Q5HQB9 | 3.40e-09 | 385 | 612 | 1128 | 1334 | Bifunctional autolysin OS=Staphylococcus epidermidis (strain ATCC 35984 / RP62A) OX=176279 GN=atl PE=3 SV=1 |
| O33635 | 3.40e-09 | 385 | 612 | 1128 | 1334 | Bifunctional autolysin OS=Staphylococcus epidermidis OX=1282 GN=atl PE=1 SV=1 |
| Q01837 | 1.30e-06 | 37 | 104 | 83 | 148 | Probable endopeptidase p60 OS=Listeria ivanovii OX=1638 GN=iap PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
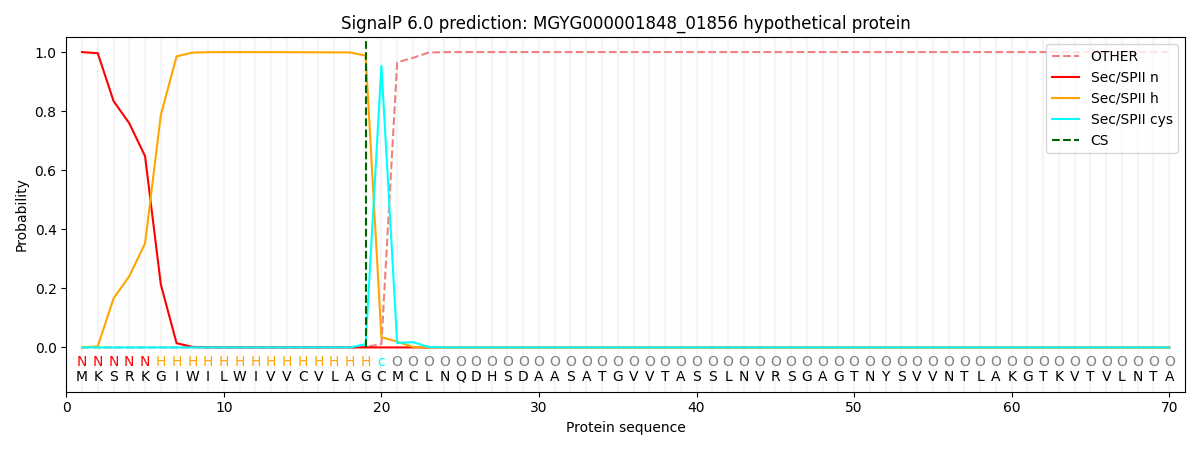
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000004 | 1.000023 | 0.000000 | 0.000000 | 0.000000 |
