You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001879_01018
You are here: Home > Sequence: MGYG000001879_01018
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; CAG-272; HGM12650; | |||||||||||
| CAZyme ID | MGYG000001879_01018 | |||||||||||
| CAZy Family | CE1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1608; End: 2657 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE1 | 39 | 238 | 2.6e-32 | 0.8898678414096917 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG4099 | COG4099 | 1.37e-46 | 30 | 259 | 160 | 387 | Predicted peptidase [General function prediction only]. |
| COG3509 | LpqC | 5.83e-22 | 20 | 212 | 28 | 208 | Poly(3-hydroxybutyrate) depolymerase [Secondary metabolites biosynthesis, transport and catabolism]. |
| COG0400 | YpfH | 1.20e-12 | 53 | 240 | 13 | 190 | Predicted esterase [General function prediction only]. |
| COG1506 | DAP2 | 3.08e-11 | 34 | 240 | 369 | 596 | Dipeptidyl aminopeptidase/acylaminoacyl peptidase [Amino acid transport and metabolism]. |
| TIGR01840 | esterase_phb | 2.59e-09 | 44 | 183 | 1 | 129 | esterase, PHB depolymerase family. This model describes a subfamily among lipases of the ab-hydrolase family. This subfamily includes bacterial depolymerases for poly(3-hydroxybutyrate) (PHB) and related polyhydroxyalkanoates (PHA), as well as acetyl xylan esterases, feruloyl esterases, and others from fungi. [Fatty acid and phospholipid metabolism, Degradation] |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BCI61582.1 | 3.38e-60 | 42 | 258 | 825 | 1042 |
| ABS60377.1 | 6.75e-42 | 24 | 258 | 4 | 244 |
| QDU56037.1 | 4.73e-40 | 32 | 258 | 788 | 1006 |
| ACR12533.1 | 4.59e-36 | 41 | 260 | 63 | 282 |
| VTR91196.1 | 2.07e-31 | 42 | 258 | 43 | 238 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3DOH_A | 1.21e-52 | 30 | 257 | 144 | 378 | CrystalStructure of a Thermostable Esterase [Thermotoga maritima],3DOH_B Crystal Structure of a Thermostable Esterase [Thermotoga maritima],3DOI_A Crystal Structure of a Thermostable Esterase complex with paraoxon [Thermotoga maritima],3DOI_B Crystal Structure of a Thermostable Esterase complex with paraoxon [Thermotoga maritima] |
| 3WYD_A | 4.38e-27 | 42 | 258 | 21 | 216 | C-terminalesterase domain of LC-Est1 [uncultured organism],3WYD_B C-terminal esterase domain of LC-Est1 [uncultured organism] |
| 4Q82_A | 6.68e-20 | 47 | 257 | 70 | 275 | CrystalStructure of Phospholipase/Carboxylesterase from Haliangium ochraceum [Haliangium ochraceum DSM 14365],4Q82_B Crystal Structure of Phospholipase/Carboxylesterase from Haliangium ochraceum [Haliangium ochraceum DSM 14365] |
| 1JJF_A | 5.71e-06 | 36 | 192 | 40 | 188 | ChainA, Endo-1,4-beta-xylanase Z [Acetivibrio thermocellus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P52090 | 2.86e-07 | 15 | 183 | 26 | 180 | Poly(3-hydroxyalkanoate) depolymerase C OS=Paucimonas lemoignei OX=29443 GN=phaZ1 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
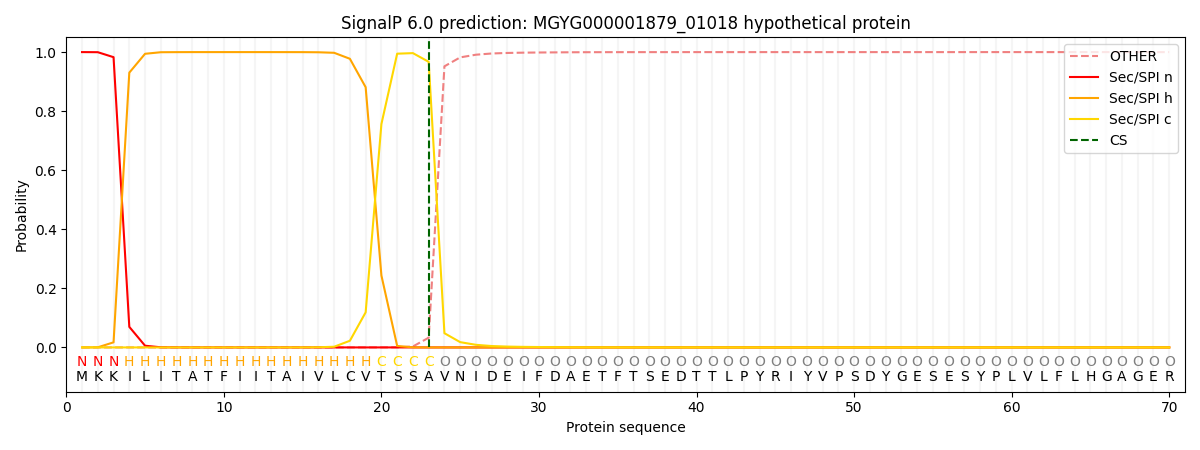
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000448 | 0.998741 | 0.000241 | 0.000192 | 0.000178 | 0.000181 |