You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001881_02114
You are here: Home > Sequence: MGYG000001881_02114
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Akkermansia muciniphila_A | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Verrucomicrobiae; Verrucomicrobiales; Akkermansiaceae; Akkermansia; Akkermansia muciniphila_A | |||||||||||
| CAZyme ID | MGYG000001881_02114 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | Beta-hexosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 5266; End: 6330 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 58 | 274 | 5.8e-44 | 0.9351851851851852 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1472 | BglX | 2.68e-74 | 1 | 309 | 5 | 311 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| PRK05337 | PRK05337 | 2.53e-63 | 1 | 309 | 1 | 305 | beta-hexosaminidase; Provisional |
| pfam00933 | Glyco_hydro_3 | 5.96e-62 | 5 | 309 | 11 | 316 | Glycosyl hydrolase family 3 N terminal domain. |
| PRK15098 | PRK15098 | 7.96e-07 | 65 | 333 | 115 | 387 | beta-glucosidase BglX. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QWP34106.1 | 5.42e-255 | 1 | 354 | 1 | 355 |
| QWP38992.1 | 5.42e-255 | 1 | 354 | 1 | 355 |
| QWP31660.1 | 5.42e-255 | 1 | 354 | 1 | 355 |
| QWP36553.1 | 5.42e-255 | 1 | 354 | 1 | 355 |
| QWP41458.1 | 5.42e-255 | 1 | 354 | 1 | 355 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7VI6_A | 7.00e-247 | 1 | 354 | 1 | 353 | ChainA, Beta-N-acetylhexosaminidase [Akkermansia muciniphila ATCC BAA-835],7VI6_B Chain B, Beta-N-acetylhexosaminidase [Akkermansia muciniphila ATCC BAA-835],7VI7_A Chain A, Beta-N-acetylhexosaminidase [Akkermansia muciniphila ATCC BAA-835],7VI7_B Chain B, Beta-N-acetylhexosaminidase [Akkermansia muciniphila ATCC BAA-835] |
| 3TEV_A | 6.94e-48 | 6 | 273 | 24 | 289 | Thecrystal structure of glycosyl hydrolase from Deinococcus radiodurans R1 [Deinococcus radiodurans R1],3TEV_B The crystal structure of glycosyl hydrolase from Deinococcus radiodurans R1 [Deinococcus radiodurans R1] |
| 5G1M_A | 6.75e-43 | 5 | 279 | 26 | 297 | Crystalstructure of NagZ from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5G1M_B Crystal structure of NagZ from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5G2M_A Crystal structure of NagZ from Pseudomonas aeruginosa in complex with N-acetylglucosamine [Pseudomonas aeruginosa PAO1],5G2M_B Crystal structure of NagZ from Pseudomonas aeruginosa in complex with N-acetylglucosamine [Pseudomonas aeruginosa PAO1],5G3R_A Crystal Structure Of Nagz From Pseudomonas Aeruginosa In Complex With N-acetylglucosamine And L-ala-1,6-anhydromurnac [Pseudomonas aeruginosa PAO1],5G3R_B Crystal Structure Of Nagz From Pseudomonas Aeruginosa In Complex With N-acetylglucosamine And L-ala-1,6-anhydromurnac [Pseudomonas aeruginosa PAO1],5G5K_A Crystal structure of NagZ from Pseudomonas aeruginosa in complex with the inhibitor 2-acetamido-1,2-dideoxynojirimycin [Pseudomonas aeruginosa PAO1],5G5K_B Crystal structure of NagZ from Pseudomonas aeruginosa in complex with the inhibitor 2-acetamido-1,2-dideoxynojirimycin [Pseudomonas aeruginosa PAO1] |
| 6JTI_A | 2.46e-42 | 3 | 313 | 46 | 357 | Crystalstructure of native NagZ from Neisseria gonorrhoeae [Neisseria gonorrhoeae],6JTI_B Crystal structure of native NagZ from Neisseria gonorrhoeae [Neisseria gonorrhoeae],6JTI_C Crystal structure of native NagZ from Neisseria gonorrhoeae [Neisseria gonorrhoeae],6JTI_D Crystal structure of native NagZ from Neisseria gonorrhoeae [Neisseria gonorrhoeae],6JTI_E Crystal structure of native NagZ from Neisseria gonorrhoeae [Neisseria gonorrhoeae],6JTI_F Crystal structure of native NagZ from Neisseria gonorrhoeae [Neisseria gonorrhoeae],6JTJ_A Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-acetylglucosamine [Neisseria gonorrhoeae],6JTJ_B Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-acetylglucosamine [Neisseria gonorrhoeae],6JTJ_C Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-acetylglucosamine [Neisseria gonorrhoeae],6JTJ_D Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-acetylglucosamine [Neisseria gonorrhoeae],6JTJ_E Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-acetylglucosamine [Neisseria gonorrhoeae],6JTJ_F Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-acetylglucosamine [Neisseria gonorrhoeae],6JTK_A Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-trifluoroacetyl-D-glucosamine [Neisseria gonorrhoeae],6JTK_B Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-trifluoroacetyl-D-glucosamine [Neisseria gonorrhoeae],6JTK_C Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-trifluoroacetyl-D-glucosamine [Neisseria gonorrhoeae],6JTK_D Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-trifluoroacetyl-D-glucosamine [Neisseria gonorrhoeae],6JTK_E Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-trifluoroacetyl-D-glucosamine [Neisseria gonorrhoeae],6JTK_F Crystal structure of NagZ from Neisseria gonorrhoeae in complex with N-trifluoroacetyl-D-glucosamine [Neisseria gonorrhoeae],6JTL_A Crystal structure of NagZ from Neisseria gonorrhoeae in complex with zinc ion [Neisseria gonorrhoeae],6JTL_B Crystal structure of NagZ from Neisseria gonorrhoeae in complex with zinc ion [Neisseria gonorrhoeae] |
| 3GS6_A | 1.48e-41 | 1 | 279 | 1 | 275 | ChainA, Beta-hexosaminidase [Vibrio cholerae],3GSM_A Chain A, Beta-hexosaminidase [Vibrio cholerae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A1SW90 | 3.28e-44 | 1 | 279 | 1 | 281 | Beta-hexosaminidase OS=Psychromonas ingrahamii (strain 37) OX=357804 GN=nagZ PE=3 SV=1 |
| Q5QUZ5 | 4.13e-43 | 1 | 279 | 1 | 279 | Beta-hexosaminidase OS=Idiomarina loihiensis (strain ATCC BAA-735 / DSM 15497 / L2-TR) OX=283942 GN=nagZ PE=3 SV=1 |
| C5BFM6 | 7.32e-43 | 1 | 296 | 1 | 294 | Beta-hexosaminidase OS=Edwardsiella ictaluri (strain 93-146) OX=634503 GN=nagZ PE=3 SV=1 |
| C4LEY6 | 1.75e-42 | 1 | 301 | 1 | 308 | Beta-hexosaminidase OS=Tolumonas auensis (strain DSM 9187 / TA4) OX=595494 GN=nagZ PE=3 SV=1 |
| Q7N397 | 2.01e-42 | 1 | 311 | 1 | 302 | Beta-hexosaminidase OS=Photorhabdus laumondii subsp. laumondii (strain DSM 15139 / CIP 105565 / TT01) OX=243265 GN=nagZ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
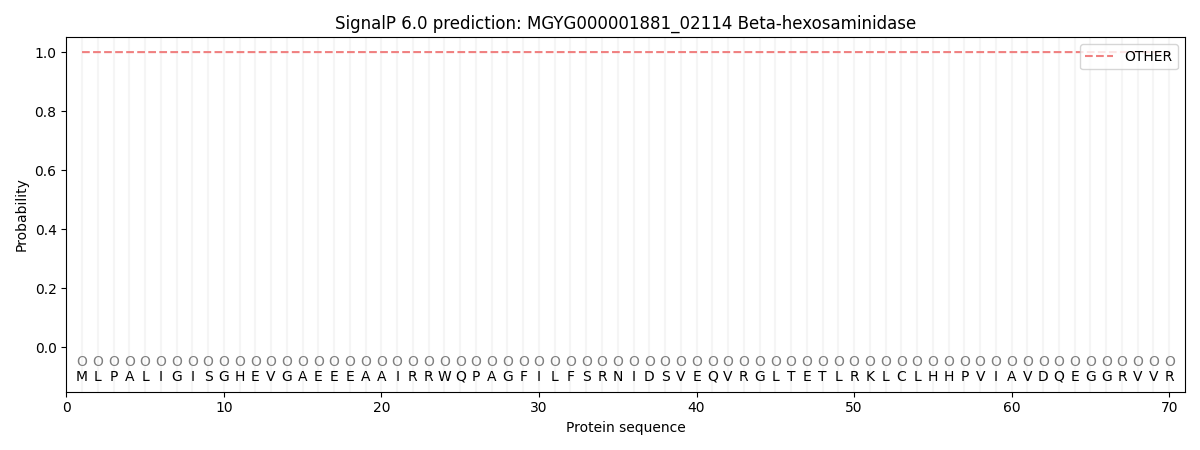
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000051 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
