You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001887_01079
You are here: Home > Sequence: MGYG000001887_01079
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Ruminiclostridium_E sp003512525 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Ruminococcaceae; Ruminiclostridium_E; Ruminiclostridium_E sp003512525 | |||||||||||
| CAZyme ID | MGYG000001887_01079 | |||||||||||
| CAZy Family | GH74 | |||||||||||
| CAZyme Description | Xyloglucanase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 47277; End: 49529 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH74 | 203 | 400 | 2.9e-99 | 0.8068669527896996 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd15482 | Sialidase_non-viral | 8.92e-12 | 86 | 354 | 32 | 321 | Non-viral sialidases. Sialidases or neuraminidases function to bind and hydrolyze terminal sialic acid residues from various glycoconjugates, they play vital roles in pathogenesis, bacterial nutrition and cellular interactions. They have a six-bladed, beta-propeller fold with the non-viral sialidases containing 2-5 Asp-box motifs (most commonly Ser/Thr-X-Asp-[X]-Gly-X-Thr- Trp/Phe). This CD includes eubacterial and eukaryotic sialidases. |
| pfam15902 | Sortilin-Vps10 | 5.13e-09 | 157 | 380 | 6 | 201 | Sortilin, neurotensin receptor 3,. Sortilin, also known in mammals as neurotensin receptor-3, is the archetypical member of a Vps10-domain (Vps10-D) that binds neurotrophic factors and neuropeptides. This domain constitutes the entire luminal part of Sortilin and is activated in the trans-Golgi network by enzymatic propeptide cleavage. The structure of the domain has been determined as a ten-bladed propeller, with up to 9 BNR or beta-hairpin turns in it. The mature receptor binds various ligands, including its own propeptide (Sort-pro), neurotensin, the pro-forms of nerve growth factor-beta (NGF)6 and brain-derived neurotrophic factor (BDNF)7, lipoprotein lipase (LpL), apo lipoprotein AV14 and the receptor-associated protein (RAP)1. |
| pfam15902 | Sortilin-Vps10 | 2.78e-08 | 562 | 711 | 1 | 167 | Sortilin, neurotensin receptor 3,. Sortilin, also known in mammals as neurotensin receptor-3, is the archetypical member of a Vps10-domain (Vps10-D) that binds neurotrophic factors and neuropeptides. This domain constitutes the entire luminal part of Sortilin and is activated in the trans-Golgi network by enzymatic propeptide cleavage. The structure of the domain has been determined as a ten-bladed propeller, with up to 9 BNR or beta-hairpin turns in it. The mature receptor binds various ligands, including its own propeptide (Sort-pro), neurotensin, the pro-forms of nerve growth factor-beta (NGF)6 and brain-derived neurotrophic factor (BDNF)7, lipoprotein lipase (LpL), apo lipoprotein AV14 and the receptor-associated protein (RAP)1. |
| pfam15902 | Sortilin-Vps10 | 5.21e-08 | 516 | 707 | 3 | 203 | Sortilin, neurotensin receptor 3,. Sortilin, also known in mammals as neurotensin receptor-3, is the archetypical member of a Vps10-domain (Vps10-D) that binds neurotrophic factors and neuropeptides. This domain constitutes the entire luminal part of Sortilin and is activated in the trans-Golgi network by enzymatic propeptide cleavage. The structure of the domain has been determined as a ten-bladed propeller, with up to 9 BNR or beta-hairpin turns in it. The mature receptor binds various ligands, including its own propeptide (Sort-pro), neurotensin, the pro-forms of nerve growth factor-beta (NGF)6 and brain-derived neurotrophic factor (BDNF)7, lipoprotein lipase (LpL), apo lipoprotein AV14 and the receptor-associated protein (RAP)1. |
| smart00602 | VPS10 | 9.41e-07 | 561 | 690 | 9 | 147 | VPS10 domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CBL33465.1 | 0.0 | 1 | 750 | 1 | 750 |
| CBK97166.1 | 0.0 | 1 | 750 | 1 | 747 |
| AHM63515.1 | 2.14e-173 | 62 | 747 | 49 | 733 |
| QPI49975.1 | 9.72e-170 | 64 | 746 | 31 | 717 |
| QEY14404.1 | 3.02e-164 | 69 | 749 | 20 | 712 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6P2L_A | 1.83e-142 | 65 | 744 | 7 | 682 | Crystalstructure of Niastella koreensis GH74 (NkGH74) enzyme [Niastella koreensis GR20-10] |
| 6P2M_A | 5.52e-112 | 64 | 746 | 10 | 691 | ChainA, Type 3a cellulose-binding domain protein [Caldicellulosiruptor lactoaceticus 6A] |
| 7KN8_A | 4.21e-93 | 69 | 747 | 15 | 713 | ChainA, Cellulase [Xanthomonas campestris pv. campestris str. ATCC 33913],7KN8_B Chain B, Cellulase [Xanthomonas campestris pv. campestris str. ATCC 33913] |
| 4LGN_A | 3.56e-90 | 64 | 748 | 7 | 735 | Thestructure of Acidothermus cellulolyticus family 74 glycoside hydrolase [Acidothermus cellulolyticus 11B] |
| 2CN3_A | 7.55e-89 | 72 | 747 | 20 | 730 | ChainA, BETA-1,4-XYLOGLUCAN HYDROLASE [Acetivibrio thermocellus],2CN3_B Chain B, BETA-1,4-XYLOGLUCAN HYDROLASE [Acetivibrio thermocellus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q3MUH7 | 1.58e-91 | 57 | 749 | 31 | 767 | Xyloglucanase OS=Paenibacillus sp. OX=58172 GN=xeg74 PE=1 SV=1 |
| Q70DK5 | 2.46e-88 | 72 | 747 | 47 | 757 | Xyloglucanase Xgh74A OS=Acetivibrio thermocellus OX=1515 GN=xghA PE=1 SV=1 |
| A3DFA0 | 4.74e-88 | 72 | 747 | 47 | 757 | Xyloglucanase Xgh74A OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=xghA PE=3 SV=1 |
| Q8J0D2 | 2.65e-84 | 64 | 750 | 27 | 787 | Oligoxyloglucan reducing end-specific cellobiohydrolase OS=Geotrichum sp. (strain M128) OX=203496 PE=1 SV=1 |
| A1DAU0 | 5.06e-80 | 59 | 749 | 22 | 799 | Probable oligoxyloglucan-reducing end-specific xyloglucanase OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=xgcA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
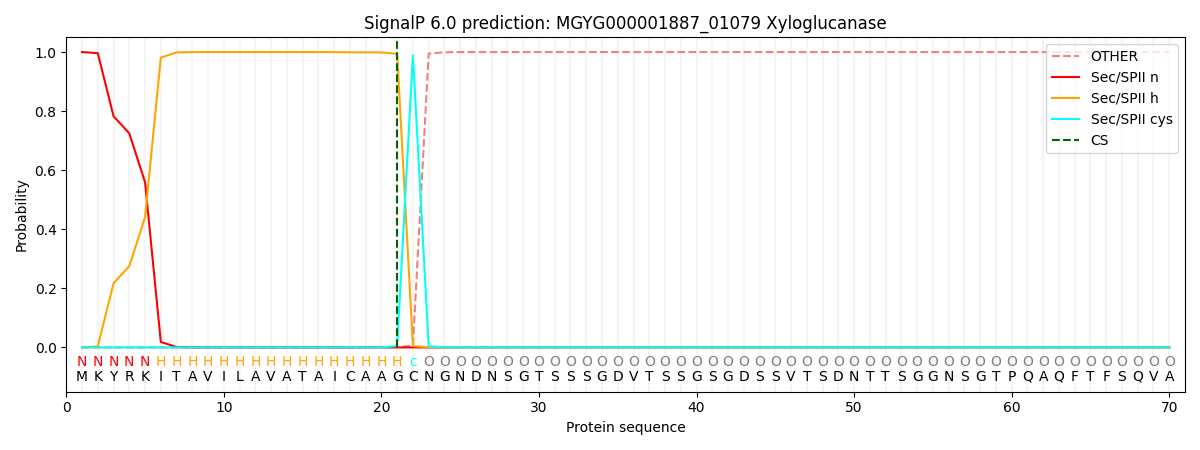
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000000 | 1.000072 | 0.000000 | 0.000000 | 0.000000 |
