You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001900_00682
You are here: Home > Sequence: MGYG000001900_00682
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Burkholderiales; Burkholderiaceae; CAG-521; | |||||||||||
| CAZyme ID | MGYG000001900_00682 | |||||||||||
| CAZy Family | CBM50 | |||||||||||
| CAZyme Description | Membrane-bound lytic murein transglycosylase D | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 58960; End: 59430 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK10783 | mltD | 8.10e-25 | 24 | 152 | 308 | 444 | membrane-bound lytic murein transglycosylase D; Provisional |
| pfam01476 | LysM | 1.68e-16 | 113 | 155 | 1 | 43 | LysM domain. The LysM (lysin motif) domain is about 40 residues long. It is found in a variety of enzymes involved in bacterial cell wall degradation. This domain may have a general peptidoglycan binding function. The structure of this domain is known. |
| pfam01476 | LysM | 7.13e-14 | 57 | 96 | 1 | 40 | LysM domain. The LysM (lysin motif) domain is about 40 residues long. It is found in a variety of enzymes involved in bacterial cell wall degradation. This domain may have a general peptidoglycan binding function. The structure of this domain is known. |
| cd00118 | LysM | 5.43e-13 | 55 | 96 | 1 | 43 | Lysin Motif is a small domain involved in binding peptidoglycan. LysM, a small globular domain with approximately 40 amino acids, is a widespread protein module involved in binding peptidoglycan in bacteria and chitin in eukaryotes. The domain was originally identified in enzymes that degrade bacterial cell walls, but proteins involved in many other biological functions also contain this domain. It has been reported that the LysM domain functions as a signal for specific plant-bacteria recognition in bacterial pathogenesis. Many of these enzymes are modular and are composed of catalytic units linked to one or several repeats of LysM domains. LysM domains are found in bacteria and eukaryotes. |
| cd00118 | LysM | 6.52e-13 | 113 | 154 | 3 | 45 | Lysin Motif is a small domain involved in binding peptidoglycan. LysM, a small globular domain with approximately 40 amino acids, is a widespread protein module involved in binding peptidoglycan in bacteria and chitin in eukaryotes. The domain was originally identified in enzymes that degrade bacterial cell walls, but proteins involved in many other biological functions also contain this domain. It has been reported that the LysM domain functions as a signal for specific plant-bacteria recognition in bacterial pathogenesis. Many of these enzymes are modular and are composed of catalytic units linked to one or several repeats of LysM domains. LysM domains are found in bacteria and eukaryotes. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QQS89247.1 | 2.39e-31 | 1 | 155 | 360 | 527 |
| BBF22593.1 | 5.86e-31 | 1 | 155 | 340 | 505 |
| QDA55156.1 | 6.53e-29 | 1 | 155 | 352 | 521 |
| ANU65188.1 | 1.31e-22 | 1 | 154 | 336 | 506 |
| QQQ96345.1 | 1.31e-22 | 1 | 154 | 336 | 506 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2MKX_A | 8.98e-06 | 57 | 96 | 7 | 46 | Solutionstructure of LysM the peptidoglycan binding domain of autolysin AtlA from Enterococcus faecalis [Enterococcus faecalis V583] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P0AEZ7 | 1.59e-13 | 24 | 154 | 305 | 442 | Membrane-bound lytic murein transglycosylase D OS=Escherichia coli (strain K12) OX=83333 GN=mltD PE=1 SV=1 |
| P0AEZ8 | 1.59e-13 | 24 | 154 | 305 | 442 | Membrane-bound lytic murein transglycosylase D OS=Escherichia coli O6:H1 (strain CFT073 / ATCC 700928 / UPEC) OX=199310 GN=mltD PE=3 SV=1 |
| P0C2T5 | 1.36e-12 | 56 | 154 | 244 | 362 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. cremoris OX=1359 GN=acmA PE=3 SV=1 |
| A2RHZ5 | 3.45e-12 | 56 | 154 | 244 | 362 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. cremoris (strain MG1363) OX=416870 GN=acmA PE=3 SV=1 |
| Q9CIT4 | 5.55e-11 | 56 | 154 | 242 | 364 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. lactis (strain IL1403) OX=272623 GN=acmA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
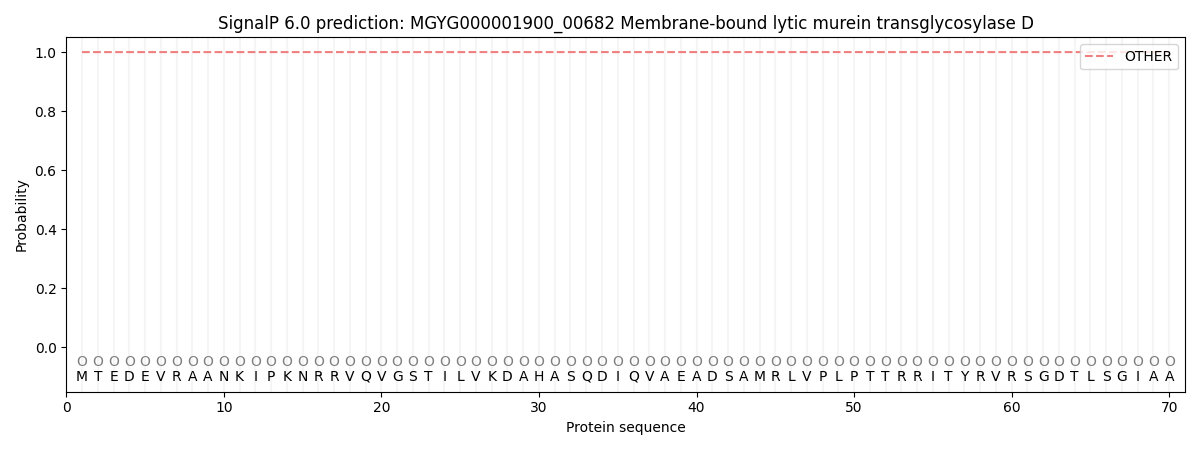
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000048 | 0.000006 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
