You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001911_01232
You are here: Home > Sequence: MGYG000001911_01232
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Desulfovibrio sp002159665 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Desulfobacterota; Desulfovibrionia; Desulfovibrionales; Desulfovibrionaceae; Desulfovibrio; Desulfovibrio sp002159665 | |||||||||||
| CAZyme ID | MGYG000001911_01232 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 19351; End: 21621 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT2 | 57 | 286 | 2.2e-28 | 0.9956521739130435 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06437 | CESA_CaSu_A2 | 3.16e-98 | 57 | 287 | 1 | 232 | Cellulose synthase catalytic subunit A2 (CESA2) is a catalytic subunit or a catalytic subunit substitute of the cellulose synthase complex. Cellulose synthase (CESA) catalyzes the polymerization reaction of cellulose using UDP-glucose as the substrate. Cellulose is an aggregate of unbranched polymers of beta-1,4-linked glucose residues, which is an abundant polysaccharide produced by plants and in varying degrees by several other organisms including algae, bacteria, fungi, and even some animals. Genomes from higher plants harbor multiple CESA genes. There are ten in Arabidopsis. At least three different CESA proteins are required to form a functional complex. In Arabidopsis, CESA1, 3 and 6 and CESA4, 7 and 8, are required for cellulose biosynthesis during primary and secondary cell wall formation. CESA2 is very closely related to CESA6 and is viewed as a prime substitute for CESA6. They functionally compensate each other. The cesa2 and cesa6 double mutant plants were significantly smaller, while the single mutant plants were almost normal. |
| cd06421 | CESA_CelA_like | 2.29e-54 | 57 | 289 | 1 | 232 | CESA_CelA_like are involved in the elongation of the glucan chain of cellulose. Family of proteins related to Agrobacterium tumefaciens CelA and Gluconacetobacter xylinus BscA. These proteins are involved in the elongation of the glucan chain of cellulose, an aggregate of unbranched polymers of beta-1,4-linked glucose residues. They are putative catalytic subunit of cellulose synthase, which is a glycosyltransferase using UDP-glucose as the substrate. The catalytic subunit is an integral membrane protein with 6 transmembrane segments and it is postulated that the protein is anchored in the membrane at the N-terminal end. |
| cd05350 | SDR_c6 | 3.62e-47 | 489 | 722 | 1 | 232 | classical (c) SDR, subgroup 6. These proteins are members of the classical SDR family, with a canonical active site tetrad and a fairly well conserved typical Gly-rich NAD-binding motif. SDRs are a functionally diverse family of oxidoreductases that have a single domain with a structurally conserved Rossmann fold (alpha/beta folding pattern with a central beta-sheet), an NAD(P)(H)-binding region, and a structurally diverse C-terminal region. Classical SDRs are typically about 250 residues long, while extended SDRS are approximately 350 residues. Sequence identity between different SDR enzymes are typically in the 15-30% range, but the enzymes share the Rossmann fold NAD-binding motif and characteristic NAD-binding and catalytic sequence patterns. These enzymes have a 3-glycine N-terminal NAD(P)(H)-binding pattern (typically, TGxxxGxG in classical SDRs and TGxxGxxG in extended SDRs), while substrate binding is in the C-terminal region. A critical catalytic Tyr residue (Tyr-151, human 15-hydroxyprostaglandin dehydrogenase (15-PGDH) numbering), is often found in a conserved YXXXK pattern. In addition to the Tyr and Lys, there is often an upstream Ser (Ser-138, 15-PGDH numbering) and/or an Asn (Asn-107, 15-PGDH numbering) or additional Ser, contributing to the active site. Substrates for these enzymes include sugars, steroids, alcohols, and aromatic compounds. The standard reaction mechanism is a proton relay involving the conserved Tyr and Lys, as well as Asn (or Ser). Some SDR family members, including 17 beta-hydroxysteroid dehydrogenase contain an additional helix-turn-helix motif that is not generally found among SDRs. |
| COG1215 | BcsA | 9.77e-44 | 1 | 396 | 15 | 403 | Glycosyltransferase, catalytic subunit of cellulose synthase and poly-beta-1,6-N-acetylglucosamine synthase [Cell motility]. |
| cd06423 | CESA_like | 5.10e-41 | 61 | 243 | 1 | 180 | CESA_like is the cellulose synthase superfamily. The cellulose synthase (CESA) superfamily includes a wide variety of glycosyltransferase family 2 enzymes that share the common characteristic of catalyzing the elongation of polysaccharide chains. The members include cellulose synthase catalytic subunit, chitin synthase, glucan biosynthesis protein and other families of CESA-like proteins. Cellulose synthase catalyzes the polymerization reaction of cellulose, an aggregate of unbranched polymers of beta-1,4-linked glucose residues in plants, most algae, some bacteria and fungi, and even some animals. In bacteria, algae and lower eukaryotes, there is a second unrelated type of cellulose synthase (Type II), which produces acylated cellulose, a derivative of cellulose. Chitin synthase catalyzes the incorporation of GlcNAc from substrate UDP-GlcNAc into chitin, which is a linear homopolymer of beta-(1,4)-linked GlcNAc residues and Glucan Biosynthesis protein catalyzes the elongation of beta-1,2 polyglucose chains of Glucan. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| SFV72799.1 | 0.0 | 1 | 735 | 1 | 729 |
| SPD36508.1 | 2.01e-303 | 1 | 736 | 1 | 738 |
| ATD80944.1 | 2.98e-303 | 1 | 736 | 12 | 749 |
| ANY15051.1 | 3.57e-131 | 4 | 482 | 8 | 487 |
| BAX59879.1 | 6.19e-129 | 7 | 486 | 10 | 491 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7LBY_A | 5.88e-21 | 52 | 289 | 268 | 503 | ChainA, Cellulose synthase catalytic subunit [UDP-forming] [Escherichia coli K-12] |
| 5EJ1_A | 5.36e-19 | 38 | 292 | 101 | 380 | ChainA, Putative cellulose synthase [Cereibacter sphaeroides 2.4.1] |
| 4HG6_A | 5.91e-19 | 38 | 292 | 113 | 392 | ChainA, Cellulose Synthase Subunit A [Cereibacter sphaeroides] |
| 4P00_A | 5.92e-19 | 38 | 292 | 114 | 393 | ChainA, Cellulose Synthase A subunit [Cereibacter sphaeroides 2.4.1],4P02_A Chain A, Cellulose Synthase subunit A [Cereibacter sphaeroides 2.4.1],5EIY_A Chain A, Putative cellulose synthase [Cereibacter sphaeroides 2.4.1],5EJZ_A Chain A, Putative cellulose synthase [Cereibacter sphaeroides 2.4.1] |
| 3AI1_A | 4.88e-16 | 490 | 672 | 11 | 193 | Thecrystal structure of L-sorbose reductase from Gluconobacter frateurii complexed with NADPH and L-sorbose reveals the structure bases of its catalytic mechanism and high substrate selectivity [Gluconobacter frateurii],3AI1_B The crystal structure of L-sorbose reductase from Gluconobacter frateurii complexed with NADPH and L-sorbose reveals the structure bases of its catalytic mechanism and high substrate selectivity [Gluconobacter frateurii],3AI2_A The crystal structure of L-sorbose reductase from Gluconobacter frateurii complexed with NADPH [Gluconobacter frateurii],3AI2_B The crystal structure of L-sorbose reductase from Gluconobacter frateurii complexed with NADPH [Gluconobacter frateurii],3AI2_C The crystal structure of L-sorbose reductase from Gluconobacter frateurii complexed with NADPH [Gluconobacter frateurii],3AI2_D The crystal structure of L-sorbose reductase from Gluconobacter frateurii complexed with NADPH [Gluconobacter frateurii],3AI2_E The crystal structure of L-sorbose reductase from Gluconobacter frateurii complexed with NADPH [Gluconobacter frateurii],3AI2_F The crystal structure of L-sorbose reductase from Gluconobacter frateurii complexed with NADPH [Gluconobacter frateurii],3AI2_G The crystal structure of L-sorbose reductase from Gluconobacter frateurii complexed with NADPH [Gluconobacter frateurii],3AI2_H The crystal structure of L-sorbose reductase from Gluconobacter frateurii complexed with NADPH [Gluconobacter frateurii] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q6YWK8 | 1.97e-39 | 57 | 286 | 118 | 353 | Probable glucomannan 4-beta-mannosyltransferase 11 OS=Oryza sativa subsp. japonica OX=39947 GN=CSLA11 PE=2 SV=1 |
| Q9FNI7 | 2.49e-38 | 57 | 286 | 97 | 332 | Glucomannan 4-beta-mannosyltransferase 2 OS=Arabidopsis thaliana OX=3702 GN=CSLA2 PE=1 SV=1 |
| Q6Z2T9 | 4.17e-38 | 53 | 286 | 137 | 376 | Probable glucomannan 4-beta-mannosyltransferase 6 OS=Oryza sativa subsp. japonica OX=39947 GN=CSLA6 PE=2 SV=2 |
| Q6UDF0 | 1.84e-37 | 43 | 312 | 76 | 353 | Glucomannan 4-beta-mannosyltransferase 1 OS=Cyamopsis tetragonoloba OX=3832 GN=ManS PE=1 SV=1 |
| Q9SRT3 | 2.12e-37 | 47 | 289 | 211 | 459 | Probable xyloglucan glycosyltransferase 6 OS=Arabidopsis thaliana OX=3702 GN=CSLC6 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
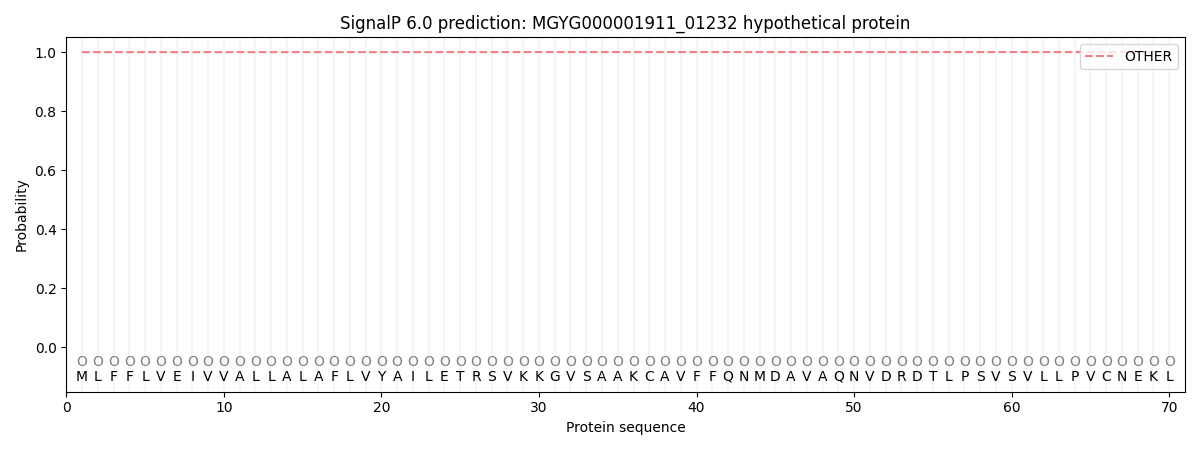
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.999793 | 0.000207 | 0.000004 | 0.000001 | 0.000001 | 0.000022 |
TMHMM Annotations download full data without filtering help
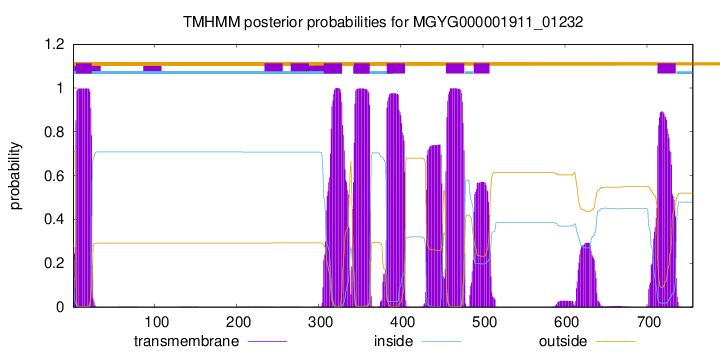
| start | end |
|---|---|
| 4 | 23 |
| 306 | 328 |
| 343 | 362 |
| 383 | 405 |
| 455 | 477 |
| 489 | 508 |
| 713 | 735 |
