You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001927_01942
You are here: Home > Sequence: MGYG000001927_01942
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp900547005 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900547005 | |||||||||||
| CAZyme ID | MGYG000001927_01942 | |||||||||||
| CAZy Family | CE3 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4014; End: 5342 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE3 | 32 | 220 | 1.1e-40 | 0.9845360824742269 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd01833 | XynB_like | 3.97e-27 | 30 | 220 | 1 | 157 | SGNH_hydrolase subfamily, similar to Ruminococcus flavefaciens XynB. Most likely a secreted hydrolase with xylanase activity. SGNH hydrolases are a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the Ser-His-Asp(Glu) triad found in other serine hydrolases. |
| pfam13472 | Lipase_GDSL_2 | 1.91e-15 | 35 | 211 | 2 | 176 | GDSL-like Lipase/Acylhydrolase family. This family of presumed lipases and related enzymes are similar to pfam00657. |
| cd00229 | SGNH_hydrolase | 5.85e-14 | 32 | 219 | 1 | 187 | SGNH_hydrolase, or GDSL_hydrolase, is a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the typical Ser-His-Asp(Glu) triad from other serine hydrolases, but may lack the carboxlic acid. |
| COG2755 | TesA | 1.86e-11 | 31 | 226 | 10 | 214 | Lysophospholipase L1 or related esterase [Amino acid transport and metabolism]. |
| cd01827 | sialate_O-acetylesterase_like1 | 4.98e-11 | 35 | 220 | 6 | 187 | sialate O-acetylesterase_like family of the SGNH hydrolases, a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the Ser-His-Asp(Glu) triad found in other serine hydrolases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BAX78799.1 | 1.46e-109 | 25 | 441 | 33 | 494 |
| ATC65549.1 | 1.60e-109 | 26 | 441 | 34 | 492 |
| QOV88633.1 | 6.94e-66 | 28 | 227 | 28 | 229 |
| QNN20991.1 | 1.34e-65 | 23 | 226 | 23 | 228 |
| QRK14019.1 | 4.58e-27 | 33 | 224 | 1 | 199 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5AO9_A | 1.71e-23 | 238 | 348 | 20 | 132 | Thestructure of a novel thermophilic esterase from the Planctomycetes species, Thermogutta terrifontis, Est2-native [Thermogutta terrifontis],5AOA_A The structure of a novel thermophilic esterase from the Planctomycetes species, Thermogutta terrifontis, Est2-Propionate bound [Thermogutta terrifontis],5AOB_A The structure of a novel thermophilic esterase from the Planctomycetes species, Thermogutta terrifontis, Est2-butyrate bound [Thermogutta terrifontis],5AOC_A The structure of a novel thermophilic esterase from the Planctomycetes species, Thermogutta terrifontis, Est2-valerate bound [Thermogutta terrifontis] |
| 7BFN_A | 1.74e-23 | 238 | 348 | 21 | 133 | ChainA, Esterase [Thermogutta terrifontis] |
| 7BFO_A | 8.37e-23 | 238 | 348 | 21 | 133 | ChainA, Esterase [Thermogutta terrifontis],7BFR_A Chain A, Esterase [Thermogutta terrifontis],7BFT_A Chain A, Esterase [Thermogutta terrifontis],7BFU_A Chain A, Esterase [Thermogutta terrifontis],7BFV_A Chain A, Esterase [Thermogutta terrifontis] |
| 2C7B_A | 1.38e-08 | 246 | 365 | 59 | 186 | TheCrystal Structure of EstE1, a New Thermophilic and Thermostable Carboxylesterase Cloned from a Metagenomic Library [uncultured archaeon],2C7B_B The Crystal Structure of EstE1, a New Thermophilic and Thermostable Carboxylesterase Cloned from a Metagenomic Library [uncultured archaeon] |
| 5B5S_A | 1.69e-08 | 32 | 213 | 4 | 192 | Crystalstructure of a carbohydrate esterase family 3 from Talaromyces cellulolyticus [Talaromyces cellulolyticus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A0A0G3FWY4 | 3.74e-10 | 249 | 351 | 62 | 162 | Probable N-octanoylanthranilate hydrolase AqdA1 OS=Rhodococcus erythropolis OX=1833 GN=aqdA1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
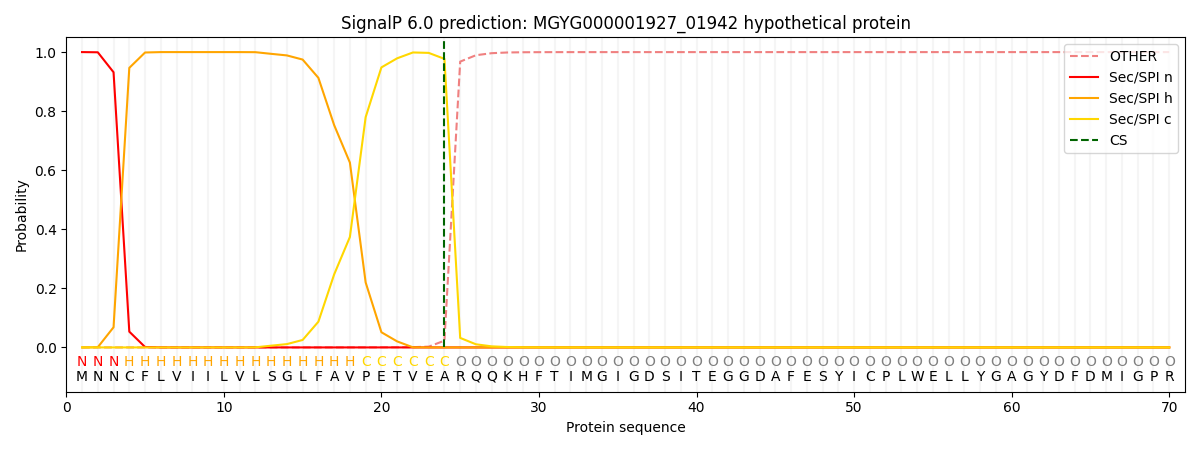
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000198 | 0.999173 | 0.000164 | 0.000161 | 0.000155 | 0.000143 |
