You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001963_02272
Basic Information
help
| Species |
Phocaeicola sp000434735
|
| Lineage |
Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Phocaeicola; Phocaeicola sp000434735
|
| CAZyme ID |
MGYG000001963_02272
|
| CAZy Family |
PL37 |
| CAZyme Description |
hypothetical protein
|
| CAZyme Property |
| Protein Length |
CGC |
Molecular Weight |
Isoelectric Point |
| 632 |
|
70738.42 |
6.4987 |
|
| Genome Property |
| Genome Assembly ID |
Genome Size |
Genome Type |
Country |
Continent |
| MGYG000001963 |
3288738 |
MAG |
Denmark |
Europe |
|
| Gene Location |
Start: 230;
End: 2128
Strand: -
|
No EC number prediction in MGYG000001963_02272.
| Family |
Start |
End |
Evalue |
family coverage |
| PL37 |
7 |
630 |
1.4e-217 |
0.9763779527559056 |
MGYG000001963_02272 has no CDD domain.
| Hit ID |
E-Value |
Query Start |
Query End |
Hit Start |
Hit End |
Description |
|
T2KMH5
|
9.95e-127 |
8 |
617 |
24 |
622 |
Broad-specificity ulvan lyase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22180 PE=1 SV=2 |
This protein is predicted as SP
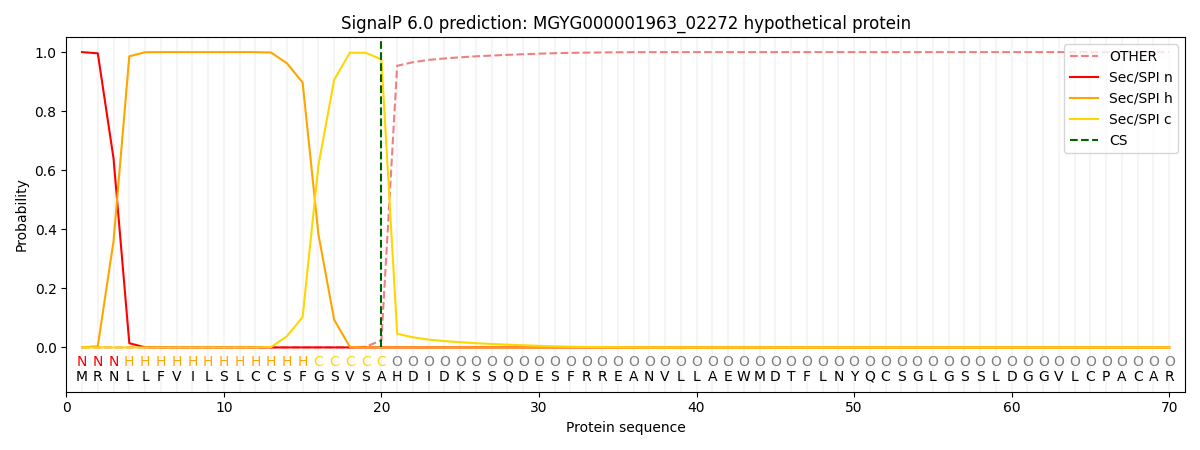
| Other |
SP_Sec_SPI |
LIPO_Sec_SPII |
TAT_Tat_SPI |
TATLIP_Sec_SPII |
PILIN_Sec_SPIII |
|
0.000268
|
0.999105
|
0.000166
|
0.000160
|
0.000151
|
0.000143
|
There is no transmembrane helices in MGYG000001963_02272.

