You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001995_00192
You are here: Home > Sequence: MGYG000001995_00192
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Parabacteroides sp900541965 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Tannerellaceae; Parabacteroides; Parabacteroides sp900541965 | |||||||||||
| CAZyme ID | MGYG000001995_00192 | |||||||||||
| CAZy Family | GT10 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 67874; End: 68806 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT10 | 113 | 270 | 7.4e-45 | 0.45821325648414984 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam18025 | FucT_N | 3.31e-40 | 6 | 97 | 1 | 92 | Alpha-(1,3)-fucosyltransferase FucT N-terminal domain. This is the N-terminal domain of the alpha chain found in Helicobacter pylori Fucosyltransferase protein which is involved in the production of Lewis x trisaccharide, a major component of lipopolysaccharide. The N-terminal domain contains the catalyst base, Glu-95 which is equivalent to the Asp-100 of other members of the glycosyltransferases-B family. The domain contains the pocket where LacNAc binds. The domain is composed of 2-10 heptad repeats and a conserved N-terminal alpha-beta-alpha motif which has little sequence similarity to the conserved N-terminal motif in other glycosyltransferases. |
| pfam00852 | Glyco_transf_10 | 4.70e-15 | 132 | 243 | 18 | 132 | Glycosyltransferase family 10 (fucosyltransferase) C-term. This is the C-terminal domain of a family of fucosyltransferases. This enzyme transfers fucose from GDP-Fucose to GlcNAc in an alpha1,3 linkage. This family is known as glycosyltransferase family 10. The C-terminal domain is the likely binding-region for ADP (manuscript in publication). |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BCI63450.1 | 1.17e-106 | 1 | 286 | 1 | 285 |
| QUT49912.1 | 1.69e-106 | 1 | 300 | 1 | 305 |
| QCT79965.1 | 9.11e-105 | 1 | 297 | 1 | 295 |
| CAH09151.1 | 1.35e-104 | 1 | 297 | 13 | 307 |
| QCQ38686.1 | 1.35e-104 | 1 | 297 | 13 | 307 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2NZW_A | 6.93e-41 | 4 | 245 | 26 | 308 | CrystalStructure of alpha1,3-Fucosyltransferase [Helicobacter pylori],2NZW_B Crystal Structure of alpha1,3-Fucosyltransferase [Helicobacter pylori],2NZW_C Crystal Structure of alpha1,3-Fucosyltransferase [Helicobacter pylori],2NZX_A Crystal Structure of alpha1,3-Fucosyltransferase with GDP [Helicobacter pylori],2NZX_B Crystal Structure of alpha1,3-Fucosyltransferase with GDP [Helicobacter pylori],2NZX_C Crystal Structure of alpha1,3-Fucosyltransferase with GDP [Helicobacter pylori],2NZY_A Crystal Structure of alpha1,3-Fucosyltransferase with GDP-fucose [Helicobacter pylori],2NZY_B Crystal Structure of alpha1,3-Fucosyltransferase with GDP-fucose [Helicobacter pylori],2NZY_C Crystal Structure of alpha1,3-Fucosyltransferase with GDP-fucose [Helicobacter pylori] |
| 5ZOI_A | 3.63e-38 | 4 | 245 | 26 | 308 | CrystalStructure of alpha1,3-Fucosyltransferase [Helicobacter pylori],5ZOI_B Crystal Structure of alpha1,3-Fucosyltransferase [Helicobacter pylori] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O30511 | 2.56e-39 | 4 | 245 | 26 | 308 | Alpha-(1,3)-fucosyltransferase FucT OS=Helicobacter pylori OX=210 GN=fucT PE=1 SV=1 |
| Q5UQ63 | 5.14e-18 | 55 | 248 | 365 | 566 | Putative fucosyltransferase R654 OS=Acanthamoeba polyphaga mimivirus OX=212035 GN=MIMI_R654 PE=3 SV=1 |
| Q6A1G3 | 2.01e-12 | 79 | 243 | 159 | 341 | Alpha-(1,3)-fucosyltransferase 10 OS=Xenopus tropicalis OX=8364 GN=fut10 PE=2 SV=1 |
| Q6A1G2 | 7.10e-12 | 132 | 245 | 264 | 391 | Alpha-(1,3)-fucosyltransferase 11 OS=Xenopus tropicalis OX=8364 GN=fut11 PE=2 SV=1 |
| Q68FV3 | 2.18e-11 | 115 | 246 | 207 | 353 | Alpha-(1,3)-fucosyltransferase 11 OS=Rattus norvegicus OX=10116 GN=Fut11 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
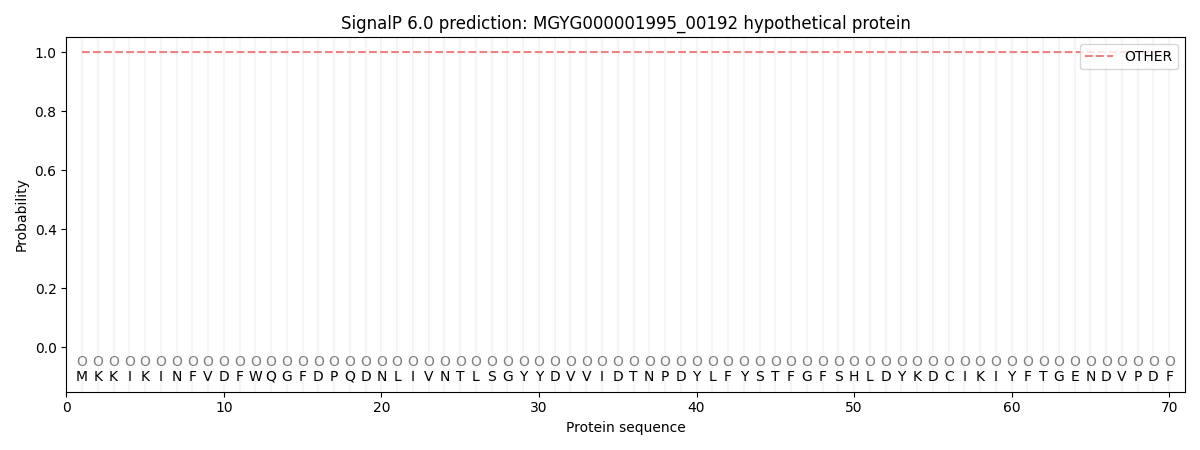
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000090 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
