You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002033_01949
You are here: Home > Sequence: MGYG000002033_01949
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Parabacteroides massiliensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Tannerellaceae; Parabacteroides; Parabacteroides massiliensis | |||||||||||
| CAZyme ID | MGYG000002033_01949 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | Beta-galactosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 3501; End: 6389 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 27 | 712 | 8.2e-69 | 0.6954787234042553 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3250 | LacZ | 6.64e-28 | 22 | 451 | 3 | 426 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| PRK10150 | PRK10150 | 5.93e-15 | 28 | 436 | 9 | 422 | beta-D-glucuronidase; Provisional |
| pfam00703 | Glyco_hydro_2 | 1.09e-11 | 204 | 308 | 3 | 106 | Glycosyl hydrolases family 2. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
| PRK10340 | ebgA | 1.11e-10 | 125 | 453 | 141 | 471 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
| pfam02837 | Glyco_hydro_2_N | 1.18e-10 | 33 | 163 | 3 | 135 | Glycosyl hydrolases family 2, sugar binding domain. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities and has a jelly-roll fold. The domain binds the sugar moiety during the sugar-hydrolysis reaction. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT50289.1 | 0.0 | 1 | 962 | 1 | 959 |
| QKH97806.1 | 0.0 | 23 | 959 | 17 | 937 |
| QUT54529.1 | 0.0 | 23 | 959 | 17 | 937 |
| QUR50384.1 | 0.0 | 23 | 959 | 17 | 937 |
| ABR43180.1 | 0.0 | 23 | 959 | 17 | 937 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6ZJV_A | 4.49e-16 | 31 | 449 | 57 | 459 | Cold-adaptedbeta-D-galactosidase from Arthrobacter sp. 32cB mutant D207A [Arthrobacter sp. 32cB],6ZJW_A Cold-adapted beta-D-galactosidase from Arthrobacter sp. 32cB mutant D207A in complex with galactose [Arthrobacter sp. 32cB],6ZJX_A Cold-adapted beta-D-galactosidase from Arthrobacter sp. 32cB mutant D207A in complex with saccharose [Arthrobacter sp. 32cB] |
| 6ETZ_A | 5.83e-16 | 31 | 449 | 36 | 438 | Cold-adaptedbeta-D-galactosidase from Arthrobacter sp. 32cB [Arthrobacter sp. 32cB],6H1P_A Cold-adapted beta-D-galactosidase from Arthrobacter sp. 32cB - data collected at room temperature [Arthrobacter sp. 32cB] |
| 6SEB_A | 5.90e-16 | 31 | 449 | 57 | 459 | Cold-adaptedbeta-D-galactosidase from Arthrobacter sp. 32cB in complex with IPTG [Arthrobacter sp. 32cB],6SEC_A Cold-adapted beta-D-galactosidase from Arthrobacter sp. 32cBon complex with ONPG [Arthrobacter sp. 32cB],6SED_A Cold-adapted beta-D-galactosidase from Arthrobacter sp. 32cB in complex with galactose [Arthrobacter sp. 32cB] |
| 6ZJP_A | 5.90e-16 | 31 | 449 | 57 | 459 | Cold-adaptedbeta-D-galactosidase from Arthrobacter sp. 32cB mutant E517Q [Arthrobacter sp. 32cB] |
| 6ZJQ_A | 5.91e-16 | 31 | 449 | 59 | 461 | Cold-adaptedbeta-D-galactosidase from Arthrobacter sp. 32cB mutant E517Q in complex with galactose [Arthrobacter sp. 32cB],6ZJR_A Cold-adapted beta-D-galactosidase from Arthrobacter sp. 32cB mutant E517Q in complex with lactulose [Arthrobacter sp. 32cB] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q59140 | 5.77e-13 | 98 | 457 | 118 | 461 | Beta-galactosidase OS=Arthrobacter sp. (strain B7) OX=86041 GN=lacZ PE=1 SV=1 |
| A4W7D2 | 6.74e-12 | 33 | 449 | 55 | 481 | Beta-galactosidase OS=Enterobacter sp. (strain 638) OX=399742 GN=lacZ PE=3 SV=1 |
| Q2XQU3 | 1.53e-11 | 30 | 449 | 52 | 481 | Beta-galactosidase 2 OS=Enterobacter cloacae OX=550 GN=lacZ PE=3 SV=1 |
| A3FEW8 | 4.00e-10 | 33 | 449 | 55 | 481 | Beta-galactosidase OS=Enterobacter agglomerans OX=549 GN=lacZ PE=1 SV=2 |
| P23989 | 9.05e-10 | 125 | 450 | 152 | 477 | Beta-galactosidase OS=Streptococcus thermophilus OX=1308 GN=lacZ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
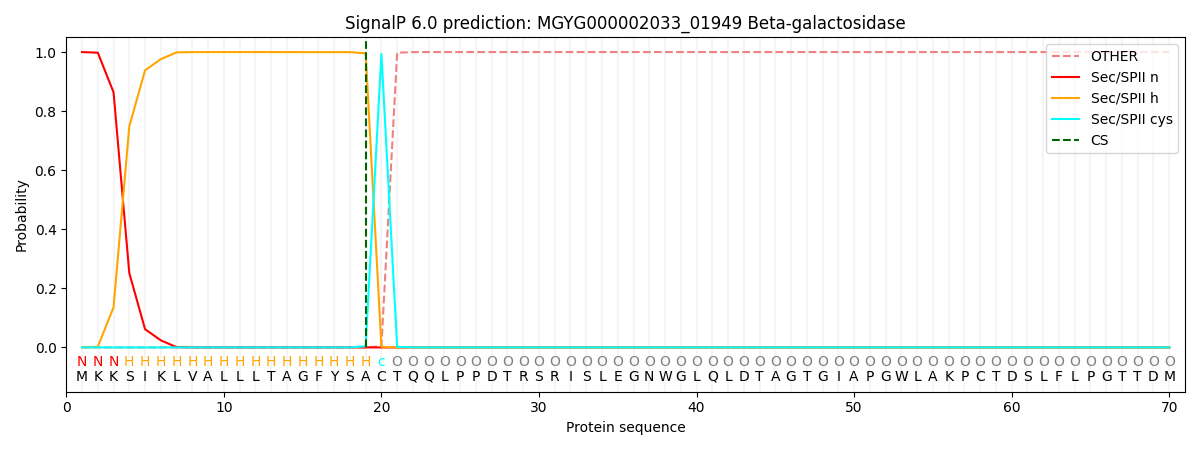
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000000 | 1.000069 | 0.000000 | 0.000000 | 0.000000 |
