You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002099_02590
You are here: Home > Sequence: MGYG000002099_02590
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia_A; Christensenellales; UBA1750; UBA7102; | |||||||||||
| CAZyme ID | MGYG000002099_02590 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 5640; End: 7262 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 71 | 329 | 9.5e-88 | 0.9883268482490273 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00150 | Cellulase | 3.48e-21 | 87 | 326 | 27 | 268 | Cellulase (glycosyl hydrolase family 5). |
| COG2730 | BglC | 1.76e-18 | 85 | 308 | 74 | 287 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| COG2723 | BglB | 3.80e-04 | 73 | 146 | 54 | 121 | Beta-glucosidase/6-phospho-beta-glucosidase/beta-galactosidase [Carbohydrate transport and metabolism]. |
| pfam13204 | DUF4038 | 0.009 | 154 | 251 | 107 | 187 | Protein of unknown function (DUF4038). A family of putative cellulases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BCK01130.1 | 8.43e-216 | 5 | 540 | 1 | 551 |
| AIQ47223.1 | 3.32e-211 | 1 | 536 | 1 | 537 |
| QSF47399.1 | 1.34e-210 | 1 | 536 | 1 | 537 |
| AIQ52770.1 | 1.09e-209 | 1 | 536 | 1 | 537 |
| BBI31985.1 | 2.36e-206 | 1 | 536 | 1 | 536 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1CEC_A | 8.46e-16 | 76 | 271 | 20 | 200 | ChainA, ENDOGLUCANASE CELC [Acetivibrio thermocellus] |
| 1CEN_A | 2.04e-15 | 76 | 271 | 20 | 200 | ChainA, CELLULASE CELC [Acetivibrio thermocellus],1CEO_A Chain A, CELLULASE CELC [Acetivibrio thermocellus] |
| 1H4P_A | 1.07e-14 | 13 | 220 | 1 | 210 | Crystalstructure of exo-1,3-beta glucanse from Saccharomyces cerevisiae [Saccharomyces cerevisiae],1H4P_B Crystal structure of exo-1,3-beta glucanse from Saccharomyces cerevisiae [Saccharomyces cerevisiae] |
| 6ZB9_A | 4.07e-14 | 31 | 285 | 3 | 275 | ChainA, Exo-beta-1,3-glucanase [uncultured bacterium],6ZB9_B Chain B, Exo-beta-1,3-glucanase [uncultured bacterium] |
| 6ZB8_A | 9.66e-14 | 31 | 285 | 3 | 275 | ChainA, Exo-beta-1,3-glucanase variant E167Q/E295Q [uncultured bacterium],6ZB8_B Chain B, Exo-beta-1,3-glucanase variant E167Q/E295Q [uncultured bacterium] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| W8QRE4 | 2.02e-25 | 13 | 309 | 3 | 283 | Beta-xylosidase OS=Phanerodontia chrysosporium OX=2822231 GN=Xyl5 PE=1 SV=2 |
| Q8NKF9 | 2.61e-18 | 31 | 237 | 37 | 240 | Glucan 1,3-beta-glucosidase OS=Candida oleophila OX=45573 GN=EXG1 PE=3 SV=1 |
| A1CRV0 | 7.77e-18 | 16 | 237 | 29 | 237 | Probable glucan 1,3-beta-glucosidase A OS=Aspergillus clavatus (strain ATCC 1007 / CBS 513.65 / DSM 816 / NCTC 3887 / NRRL 1 / QM 1276 / 107) OX=344612 GN=exgA PE=3 SV=2 |
| A3DJ77 | 3.45e-15 | 76 | 271 | 20 | 200 | Endoglucanase C OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celC PE=3 SV=1 |
| P23340 | 3.45e-15 | 76 | 271 | 20 | 200 | Endoglucanase C307 OS=Clostridium sp. (strain F1) OX=1508 GN=celC307 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
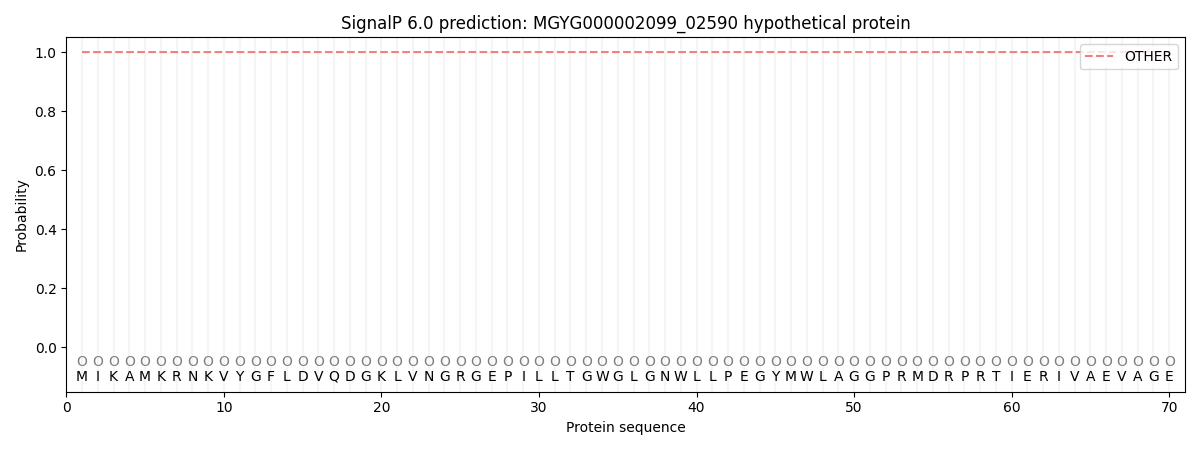
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000061 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
