You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002108_00724
You are here: Home > Sequence: MGYG000002108_00724
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
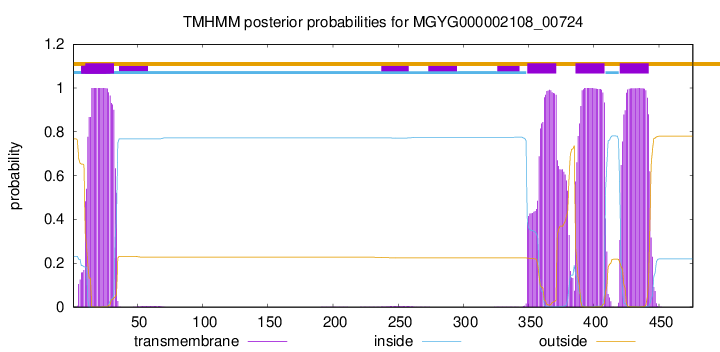
TMHMM annotations
Basic Information help
| Species | Prevotella sp900290275 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900290275 | |||||||||||
| CAZyme ID | MGYG000002108_00724 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 36583; End: 38013 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT2 | 62 | 209 | 1.6e-20 | 0.7235294117647059 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06423 | CESA_like | 1.25e-54 | 63 | 280 | 1 | 180 | CESA_like is the cellulose synthase superfamily. The cellulose synthase (CESA) superfamily includes a wide variety of glycosyltransferase family 2 enzymes that share the common characteristic of catalyzing the elongation of polysaccharide chains. The members include cellulose synthase catalytic subunit, chitin synthase, glucan biosynthesis protein and other families of CESA-like proteins. Cellulose synthase catalyzes the polymerization reaction of cellulose, an aggregate of unbranched polymers of beta-1,4-linked glucose residues in plants, most algae, some bacteria and fungi, and even some animals. In bacteria, algae and lower eukaryotes, there is a second unrelated type of cellulose synthase (Type II), which produces acylated cellulose, a derivative of cellulose. Chitin synthase catalyzes the incorporation of GlcNAc from substrate UDP-GlcNAc into chitin, which is a linear homopolymer of beta-(1,4)-linked GlcNAc residues and Glucan Biosynthesis protein catalyzes the elongation of beta-1,2 polyglucose chains of Glucan. |
| COG1215 | BcsA | 3.72e-46 | 1 | 471 | 1 | 431 | Glycosyltransferase, catalytic subunit of cellulose synthase and poly-beta-1,6-N-acetylglucosamine synthase [Cell motility]. |
| PRK11204 | PRK11204 | 3.17e-32 | 5 | 423 | 5 | 378 | N-glycosyltransferase; Provisional |
| PRK14583 | hmsR | 2.46e-21 | 13 | 398 | 33 | 383 | poly-beta-1,6 N-acetyl-D-glucosamine synthase. |
| cd02525 | Succinoglycan_BP_ExoA | 1.16e-20 | 61 | 349 | 2 | 243 | ExoA is involved in the biosynthesis of succinoglycan. Succinoglycan Biosynthesis Protein ExoA catalyzes the formation of a beta-1,3 linkage of the second sugar (glucose) of the succinoglycan with the galactose on the lipid carrie. Succinoglycan is an acidic exopolysaccharide that is important for invasion of the nodules. Succinoglycan is a high-molecular-weight polymer composed of repeating octasaccharide units. These units are synthesized on membrane-bound isoprenoid lipid carriers, beginning with galactose followed by seven glucose molecules, and modified by the addition of acetate, succinate, and pyruvate. ExoA is a membrane protein with a transmembrance domain at c-terminus. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT74207.1 | 5.82e-195 | 3 | 474 | 2 | 476 |
| QIU93724.1 | 6.07e-193 | 3 | 470 | 2 | 472 |
| CBK67631.1 | 3.88e-191 | 10 | 470 | 9 | 472 |
| QUT31986.1 | 3.88e-191 | 10 | 470 | 9 | 472 |
| QUT26577.1 | 3.88e-191 | 10 | 470 | 9 | 472 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6P61_A | 6.43e-07 | 59 | 102 | 13 | 58 | Structureof a Glycosyltransferase from Leptospira borgpetersenii serovar Hardjo-bovis (strain JB197) [Leptospira borgpetersenii serovar Hardjo-bovis str. JB197],6P61_B Structure of a Glycosyltransferase from Leptospira borgpetersenii serovar Hardjo-bovis (strain JB197) [Leptospira borgpetersenii serovar Hardjo-bovis str. JB197],6P61_C Structure of a Glycosyltransferase from Leptospira borgpetersenii serovar Hardjo-bovis (strain JB197) [Leptospira borgpetersenii serovar Hardjo-bovis str. JB197],6P61_D Structure of a Glycosyltransferase from Leptospira borgpetersenii serovar Hardjo-bovis (strain JB197) [Leptospira borgpetersenii serovar Hardjo-bovis str. JB197] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q5HKQ0 | 1.77e-17 | 60 | 329 | 48 | 270 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus epidermidis (strain ATCC 35984 / RP62A) OX=176279 GN=icaA PE=1 SV=1 |
| Q8GLC5 | 3.18e-17 | 60 | 329 | 48 | 270 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus epidermidis OX=1282 GN=icaA PE=3 SV=1 |
| Q9RQP9 | 3.38e-15 | 60 | 329 | 48 | 270 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus aureus (strain NCTC 8325 / PS 47) OX=93061 GN=icaA PE=3 SV=2 |
| Q99QX3 | 3.38e-15 | 60 | 329 | 48 | 270 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus aureus (strain Mu50 / ATCC 700699) OX=158878 GN=icaA PE=3 SV=1 |
| Q5HCN1 | 3.38e-15 | 60 | 329 | 48 | 270 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus aureus (strain COL) OX=93062 GN=icaA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
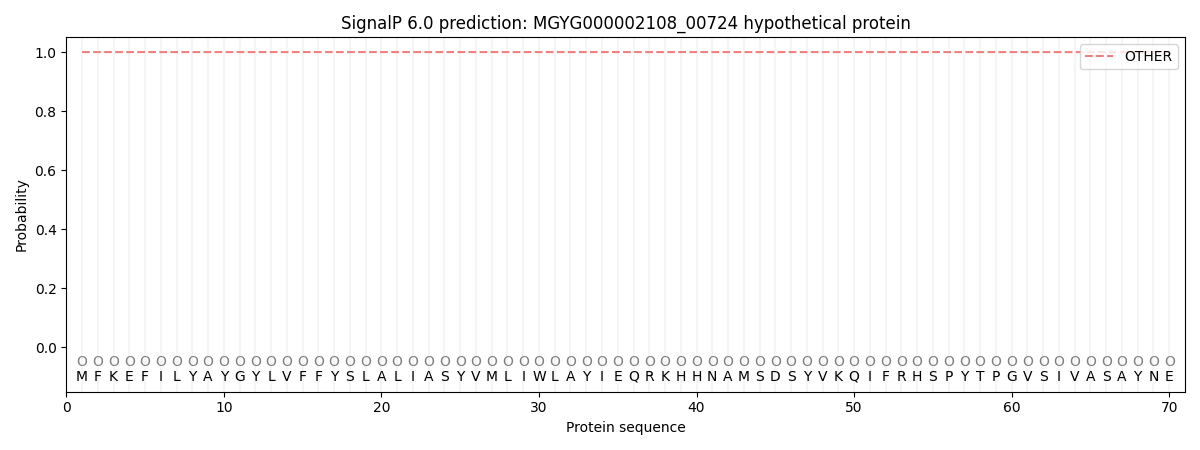
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000058 | 0.000001 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |