You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002131_01144
Basic Information
help
| Species |
CAG-882 sp900545455
|
| Lineage |
Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; CAG-882; CAG-882 sp900545455
|
| CAZyme ID |
MGYG000002131_01144
|
| CAZy Family |
GT2 |
| CAZyme Description |
hypothetical protein
|
| CAZyme Property |
|
| Genome Property |
| Genome Assembly ID |
Genome Size |
Genome Type |
Country |
Continent |
| MGYG000002131 |
2861336 |
MAG |
United States |
North America |
|
| Gene Location |
Start: 4193;
End: 5149
Strand: +
|
No EC number prediction in MGYG000002131_01144.
MGYG000002131_01144 has no CDD domain.
| Hit ID |
E-Value |
Query Start |
Query End |
Hit Start |
Hit End |
| AOZ97351.1
|
3.04e-114 |
5 |
316 |
4 |
317 |
| AOZ97352.1
|
5.06e-112 |
5 |
316 |
321 |
634 |
| ADL33240.1
|
1.17e-111 |
3 |
316 |
2 |
316 |
This protein is predicted as OTHER
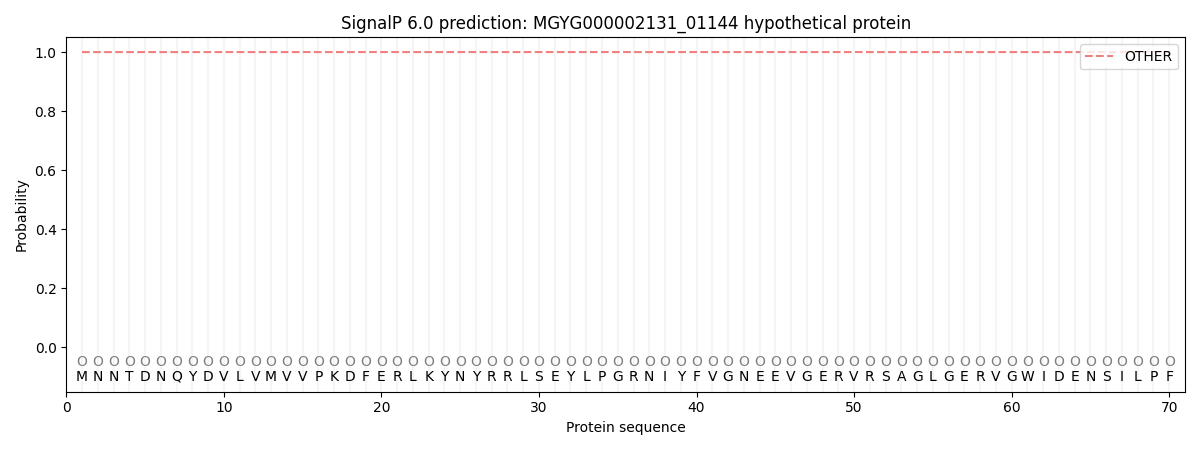
| Other |
SP_Sec_SPI |
LIPO_Sec_SPII |
TAT_Tat_SPI |
TATLIP_Sec_SPII |
PILIN_Sec_SPIII |
|
1.000074
|
0.000000
|
0.000000
|
0.000000
|
0.000000
|
0.000000
|
There is no transmembrane helices in MGYG000002131_01144.

