You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002148_02828
You are here: Home > Sequence: MGYG000002148_02828
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Eisenbergiella; | |||||||||||
| CAZyme ID | MGYG000002148_02828 | |||||||||||
| CAZy Family | GH25 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 39376; End: 40791 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH25 | 38 | 206 | 4.3e-45 | 0.9943502824858758 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06414 | GH25_LytC-like | 5.07e-64 | 36 | 216 | 2 | 191 | The LytC lysozyme of Streptococcus pneumoniae is a bacterial cell wall hydrolase that cleaves the beta1-4-glycosydic bond located between the N-acetylmuramoyl-N-glucosaminyl residues of the cell wall polysaccharide chains. LytC is composed of a C-terminal glycosyl hydrolase family 25 (GH25) domain and an N-terminal choline-binding module (CBM) consisting of eleven homologous repeats that specifically recognizes the choline residues of pneumococcal lipoteichoic and teichoic acids. This domain arrangement is the reverse of the major pneumococcal autolysin, LytA, and the CPL-1-like lytic enzymes of the pneumococcal bacteriophages, in which the CBM (consisting of six repeats) is at the C-terminus. This model represents the C-terminal catalytic domain of the LytC-like enzymes. |
| cd00599 | GH25_muramidase | 9.38e-34 | 36 | 214 | 1 | 185 | Endo-N-acetylmuramidases (muramidases) are lysozymes (also referred to as peptidoglycan hydrolases) that degrade bacterial cell walls by catalyzing the hydrolysis of 1,4-beta-linkages between N-acetylmuramic acid and N-acetyl-D-glucosamine residues. This family of muramidases contains a glycosyl hydrolase family 25 (GH25) catalytic domain and is found in bacteria, fungi, slime molds, round worms, protozoans and bacteriophages. The bacteriophage members are referred to as endolysins which are involved in lysing the host cell at the end of the replication cycle to allow release of mature phage particles. Endolysins are typically modular enzymes consisting of a catalytically active domain that hydrolyzes the peptidoglycan cell wall and a cell wall-binding domain that anchors the protein to the cell wall. Endolysins generally have narrow substrate specificities with either intra-species or intra-genus bacteriolytic activity. |
| pfam01183 | Glyco_hydro_25 | 7.24e-27 | 38 | 206 | 1 | 180 | Glycosyl hydrolases family 25. |
| NF033930 | pneumo_PspA | 2.89e-26 | 225 | 426 | 462 | 657 | pneumococcal surface protein A. The pneumococcal surface protein proteins, found in Streptococcus pneumoniae, are repetitive, with patterns of localized high sequence identity across pairs of proteins given different specific names that recombination may be presumed. This protein, PspA, has an N-terminal region that lacks a cross-wall-targeting YSIRK type extended signal peptide, in contrast to the closely related choline-binding protein CbpA which has a similar C-terminus but a YSIRK-containing region at the N-terminus. |
| NF033838 | PspC_subgroup_1 | 1.34e-23 | 246 | 426 | 487 | 680 | pneumococcal surface protein PspC, choline-binding form. The pneumococcal surface protein PspC, as described in Streptococcus pneumoniae, is a repetitive and highly variable protein, recognized by a conserved N-terminal domain and also by genomic location. This form, subgroup 1, has variable numbers of a choline-binding repeat in the C-terminal region, and is also known as choline-binding protein A. The other form, subgroup 2, is anchored covalently after cleavage by sortase at a C-terminal LPXTG site. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ADL32940.1 | 2.24e-67 | 34 | 304 | 39 | 310 |
| AOZ95227.1 | 9.20e-62 | 2 | 312 | 3 | 319 |
| CBK77005.1 | 3.31e-53 | 35 | 285 | 51 | 301 |
| QQR03194.1 | 5.57e-50 | 34 | 237 | 99 | 303 |
| ANU47907.1 | 5.57e-50 | 34 | 237 | 99 | 303 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2WAG_A | 1.72e-13 | 36 | 213 | 17 | 204 | TheStructure of a family 25 Glycosyl hydrolase from Bacillus anthracis. [Bacillus anthracis str. Ames] |
| 2WW5_A | 5.42e-12 | 36 | 213 | 270 | 465 | 3D-structureof the modular autolysin LytC from Streptococcus pneumoniae at 1.6 A resolution [Streptococcus pneumoniae R6],2WWD_A 3D-structure of the modular autolysin LytC from Streptococcus pneumoniae in complex with pneummococcal peptidoglycan fragment [Streptococcus pneumoniae R6] |
| 2WWC_A | 1.27e-11 | 36 | 213 | 270 | 465 | 3D-structureof the modular autolysin LytC from Streptococcus pneumoniae in complex with synthetic peptidoglycan ligand [Streptococcus pneumoniae R6] |
| 1JFX_A | 2.08e-11 | 36 | 213 | 6 | 200 | Crystalstructure of the bacterial lysozyme from Streptomyces coelicolor at 1.65 A resolution [Streptomyces coelicolor] |
| 4KRU_A | 5.96e-08 | 36 | 213 | 21 | 205 | X-raystructure of catalytic domain of endolysin from clostridium perfringens phage phiSM101 [Clostridium phage phiSM101] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P25310 | 2.91e-10 | 36 | 213 | 83 | 277 | Lysozyme M1 OS=Streptomyces globisporus OX=1908 GN=acm PE=1 SV=1 |
| P26836 | 9.72e-07 | 36 | 228 | 10 | 207 | Probable autolytic lysozyme OS=Clostridium perfringens (strain 13 / Type A) OX=195102 GN=lyc PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
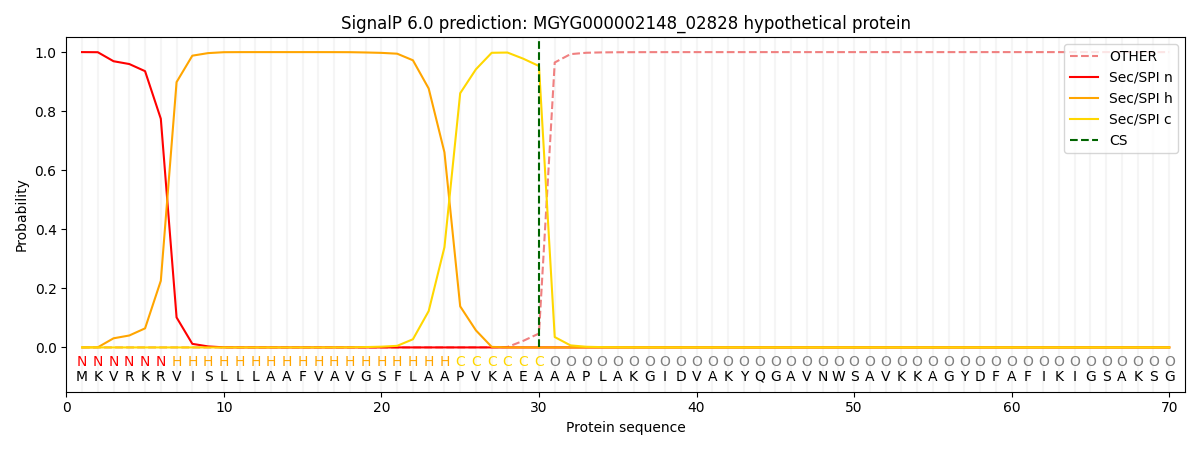
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000233 | 0.999123 | 0.000152 | 0.000190 | 0.000156 | 0.000140 |

