You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002176_00621
You are here: Home > Sequence: MGYG000002176_00621
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
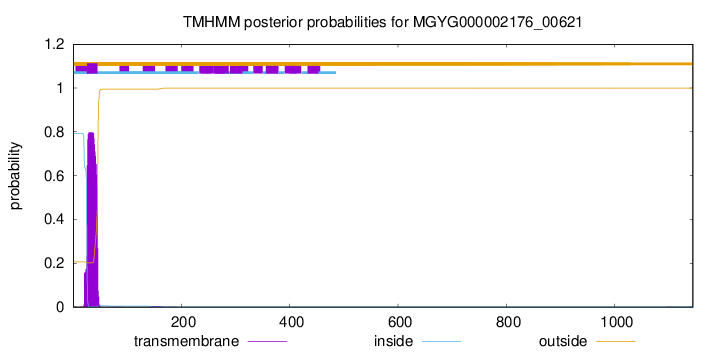
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Muribaculaceae; SFTJ01; | |||||||||||
| CAZyme ID | MGYG000002176_00621 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 10004; End: 13438 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 89 | 328 | 2.1e-56 | 0.9814814814814815 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam16147 | DUF4855 | 3.27e-127 | 821 | 1130 | 1 | 313 | Domain of unknown function (DUF4855). This family consists of uncharacterized proteins around 400 residues in length and is mainly found in various Bacteroides species. Several proteins are annotated as glycerophosphodiester phosphodiesterases, but the origin of this annotation is not clear. |
| PRK15098 | PRK15098 | 2.86e-78 | 146 | 747 | 118 | 730 | beta-glucosidase BglX. |
| COG1472 | BglX | 4.87e-56 | 142 | 426 | 77 | 370 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| pfam01915 | Glyco_hydro_3_C | 2.70e-46 | 394 | 666 | 1 | 216 | Glycosyl hydrolase family 3 C-terminal domain. This domain is involved in catalysis and may be involved in binding beta-glucan. This domain is found associated with pfam00933. |
| PLN03080 | PLN03080 | 1.51e-33 | 55 | 738 | 45 | 739 | Probable beta-xylosidase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT74321.1 | 1.27e-304 | 49 | 807 | 33 | 766 |
| AHF25042.1 | 1.95e-214 | 47 | 806 | 25 | 776 |
| AFL77429.1 | 2.98e-212 | 33 | 807 | 14 | 775 |
| CBK63831.1 | 4.21e-212 | 33 | 807 | 14 | 775 |
| AFX98011.1 | 2.59e-211 | 34 | 804 | 12 | 774 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2X40_A | 2.56e-175 | 50 | 792 | 4 | 720 | Structureof beta-glucosidase 3B from Thermotoga neapolitana in complex with glycerol [Thermotoga neapolitana DSM 4359],2X41_A Structure of beta-glucosidase 3B from Thermotoga neapolitana in complex with glucose [Thermotoga neapolitana DSM 4359] |
| 2X42_A | 3.93e-174 | 50 | 792 | 4 | 720 | Structureof beta-glucosidase 3B from Thermotoga neapolitana in complex with alpha-D-glucose [Thermotoga neapolitana DSM 4359] |
| 7MS2_A | 1.50e-107 | 49 | 772 | 4 | 657 | ChainA, Thermostable beta-glucosidase B [Acetivibrio thermocellus AD2],7MS2_B Chain B, Thermostable beta-glucosidase B [Acetivibrio thermocellus AD2] |
| 5WAB_A | 9.43e-105 | 54 | 772 | 9 | 652 | CrystalStructure of Bifidobacterium adolescentis GH3 beta-glucosidase [Bifidobacterium adolescentis ATCC 15703],5WAB_B Crystal Structure of Bifidobacterium adolescentis GH3 beta-glucosidase [Bifidobacterium adolescentis ATCC 15703],5WAB_C Crystal Structure of Bifidobacterium adolescentis GH3 beta-glucosidase [Bifidobacterium adolescentis ATCC 15703],5WAB_D Crystal Structure of Bifidobacterium adolescentis GH3 beta-glucosidase [Bifidobacterium adolescentis ATCC 15703] |
| 3AC0_A | 9.07e-85 | 87 | 792 | 30 | 843 | Crystalstructure of Beta-glucosidase from Kluyveromyces marxianus in complex with glucose [Kluyveromyces marxianus],3AC0_B Crystal structure of Beta-glucosidase from Kluyveromyces marxianus in complex with glucose [Kluyveromyces marxianus],3AC0_C Crystal structure of Beta-glucosidase from Kluyveromyces marxianus in complex with glucose [Kluyveromyces marxianus],3AC0_D Crystal structure of Beta-glucosidase from Kluyveromyces marxianus in complex with glucose [Kluyveromyces marxianus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P14002 | 8.23e-107 | 49 | 772 | 4 | 657 | Thermostable beta-glucosidase B OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=bglB PE=1 SV=2 |
| E7CY69 | 5.74e-102 | 54 | 768 | 9 | 657 | Exo-alpha-(1->6)-L-arabinopyranosidase OS=Bifidobacterium longum OX=216816 GN=apy PE=1 SV=1 |
| F6C6C1 | 5.60e-101 | 54 | 772 | 9 | 661 | Exo-alpha-(1->6)-L-arabinopyranosidase OS=Bifidobacterium breve (strain ACS-071-V-Sch8b) OX=866777 GN=HMPREF9228_1477 PE=1 SV=1 |
| Q5BFG8 | 6.55e-96 | 50 | 771 | 11 | 826 | Beta-glucosidase B OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=bglB PE=1 SV=1 |
| B8NDE2 | 1.90e-85 | 87 | 791 | 30 | 836 | Probable beta-glucosidase I OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=bglI PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
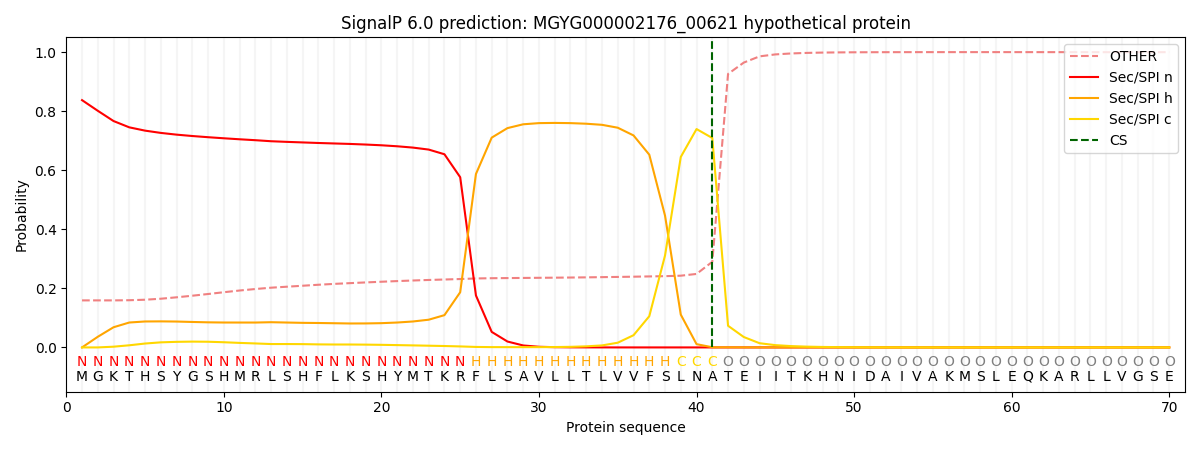
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.163179 | 0.831711 | 0.004250 | 0.000293 | 0.000259 | 0.000291 |