You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002190_01197
You are here: Home > Sequence: MGYG000002190_01197
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Clostridium sp001916075 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Clostridiales; Clostridiaceae; Clostridium; Clostridium sp001916075 | |||||||||||
| CAZyme ID | MGYG000002190_01197 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | Beta-glucanase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 68778; End: 69503 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH16 | 36 | 236 | 3e-92 | 0.9757281553398058 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd02175 | GH16_lichenase | 3.54e-121 | 31 | 240 | 1 | 212 | lichenase, member of glycosyl hydrolase family 16. Lichenase, also known as 1,3-1,4-beta-glucanase, is a member of glycosyl hydrolase family 16, that specifically cleaves 1,4-beta-D-glucosidic bonds in mixed-linked beta glucans that also contain 1,3-beta-D-glucosidic linkages. Natural substrates of beta-glucanase are beta-glucans from grain endosperm cell walls or lichenan from the Islandic moss, Cetraria islandica. This protein is found not only in bacteria but also in anaerobic fungi. This domain includes two seven-stranded antiparallel beta-sheets that are adjacent to one another forming a compact, jellyroll beta-sandwich structure. |
| COG2273 | BglS | 2.07e-57 | 31 | 238 | 47 | 264 | Beta-glucanase, GH16 family [Carbohydrate transport and metabolism]. |
| pfam00722 | Glyco_hydro_16 | 5.28e-53 | 59 | 236 | 1 | 167 | Glycosyl hydrolases family 16. |
| cd00413 | Glyco_hydrolase_16 | 8.80e-37 | 42 | 236 | 12 | 207 | glycosyl hydrolase family 16. The O-Glycosyl hydrolases are a widespread group of enzymes that hydrolyse the glycosidic bond between two or more carbohydrates, or between a carbohydrate and a non-carbohydrate moiety. A glycosyl hydrolase classification system based on sequence similarity has led to the definition of more than 95 different families inlcuding glycosyl hydrolase family 16. Family 16 includes lichenase, xyloglucan endotransglycosylase (XET), beta-agarase, kappa-carrageenase, endo-beta-1,3-glucanase, endo-beta-1,3-1,4-glucanase, and endo-beta-galactosidase, all of which have a conserved jelly roll fold with a deep active site channel harboring the catalytic residues. |
| cd02176 | GH16_XET | 9.49e-27 | 55 | 190 | 10 | 142 | Xyloglucan endotransglycosylase, member of glycosyl hydrolase family 16. Xyloglucan endotransglycosylases (XETs) cleave and religate xyloglucan polymers in plant cell walls via a transglycosylation mechanism. Xyloglucan is a soluble hemicellulose with a backbone of beta-1,4-linked glucose units, partially substituted with alpha-1,6-linked xylopyranose branches. It binds noncovalently to cellulose, cross-linking the adjacent cellulose microfibrils, giving it a key structural role as a matrix polymer. Therefore, XET plays an important role in all plant processes that require cell wall remodeling. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AGF58046.1 | 2.46e-136 | 1 | 241 | 6 | 246 |
| AQR96736.1 | 2.86e-135 | 1 | 241 | 6 | 246 |
| ADL52419.1 | 2.97e-135 | 1 | 241 | 1 | 247 |
| BCZ45550.1 | 3.88e-133 | 1 | 240 | 6 | 245 |
| BCZ47763.1 | 5.71e-133 | 11 | 241 | 16 | 246 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3I4I_A | 8.14e-115 | 29 | 241 | 22 | 234 | Crystalstructure of a prokaryotic beta-1,3-1,4-glucanase (lichenase) derived from a mouse hindgut metagenome [uncultured murine large bowel bacterium BAC 14],3I4I_B Crystal structure of a prokaryotic beta-1,3-1,4-glucanase (lichenase) derived from a mouse hindgut metagenome [uncultured murine large bowel bacterium BAC 14] |
| 1MAC_A | 3.93e-96 | 29 | 238 | 2 | 209 | CrystalStructure And Site-Directed Mutagenesis Of Bacillus Macerans Endo-1,3-1,4-Beta-Glucanase [Paenibacillus macerans],1MAC_B Crystal Structure And Site-Directed Mutagenesis Of Bacillus Macerans Endo-1,3-1,4-Beta-Glucanase [Paenibacillus macerans] |
| 3O5S_A | 1.34e-95 | 20 | 238 | 19 | 235 | CrystalStructure of the endo-beta-1,3-1,4 glucanase from Bacillus subtilis (strain 168) [Bacillus subtilis] |
| 3WVJ_A | 1.71e-94 | 36 | 240 | 13 | 215 | ChainA, Beta-glucanase [Acetivibrio thermocellus ATCC 27405],3WVJ_B Chain B, Beta-glucanase [Acetivibrio thermocellus ATCC 27405] |
| 1BYH_A | 3.98e-94 | 27 | 238 | 2 | 211 | MOLECULARAND ACTIVE-SITE STRUCTURE OF A BACILLUS (1-3,1-4)-BETA-GLUCANASE [synthetic construct],1GLH_A Cation Binding To A Bacillus (1,3-1,4)-Beta-Glucanase. Geometry, Affinity And Effect On Protein Stability [Paenibacillus macerans],2AYH_A Crystal And Molecular Structure At 1.6 Angstroms Resolution Of The Hybrid Bacillus Endo-1,3-1,4-Beta-D-Glucan 4- Glucanohydrolase H(A16-M) [hybrid] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P45797 | 1.66e-98 | 23 | 238 | 22 | 235 | Beta-glucanase OS=Paenibacillus polymyxa OX=1406 GN=gluB PE=3 SV=1 |
| P23904 | 3.74e-97 | 23 | 238 | 21 | 234 | Beta-glucanase OS=Paenibacillus macerans OX=44252 PE=1 SV=2 |
| P04957 | 1.80e-96 | 8 | 238 | 11 | 239 | Beta-glucanase OS=Bacillus subtilis (strain 168) OX=224308 GN=bglS PE=1 SV=2 |
| P07980 | 3.78e-95 | 1 | 238 | 1 | 236 | Beta-glucanase OS=Bacillus amyloliquefaciens OX=1390 GN=bglA PE=3 SV=1 |
| A3DBX3 | 3.96e-92 | 36 | 240 | 42 | 244 | Beta-glucanase OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=licB PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
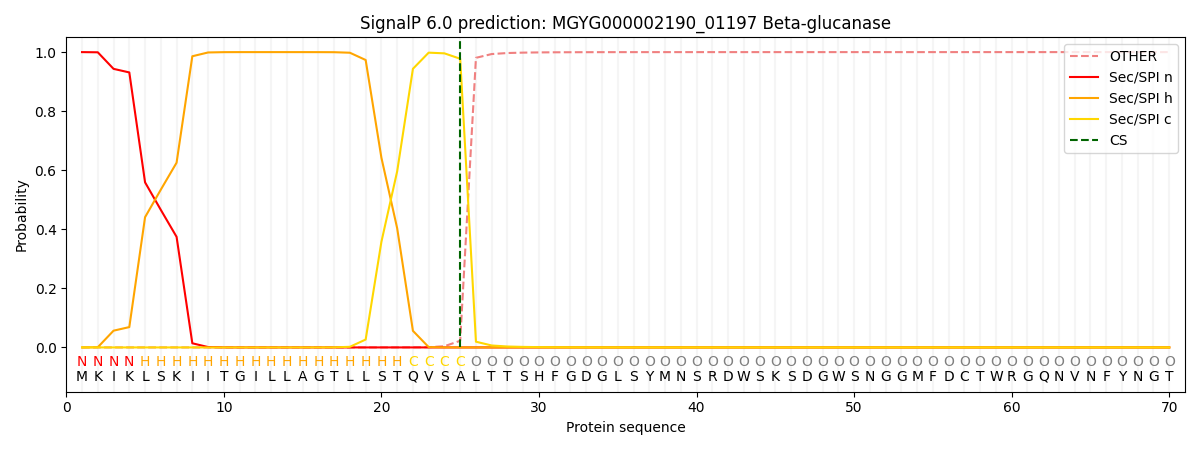
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000218 | 0.999090 | 0.000183 | 0.000179 | 0.000159 | 0.000150 |
