You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002258_01254
You are here: Home > Sequence: MGYG000002258_01254
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UBA1081 sp900543395 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Acutalibacteraceae; UBA1081; UBA1081 sp900543395 | |||||||||||
| CAZyme ID | MGYG000002258_01254 | |||||||||||
| CAZy Family | GH78 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 9558; End: 11078 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH78 | 103 | 442 | 2.8e-22 | 0.5634920634920635 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3408 | GDB1 | 2.28e-16 | 63 | 445 | 256 | 620 | Glycogen debranching enzyme (alpha-1,6-glucosidase) [Carbohydrate transport and metabolism]. |
| PRK10137 | PRK10137 | 1.96e-13 | 146 | 403 | 417 | 758 | alpha-glucosidase; Provisional |
| pfam01204 | Trehalase | 8.41e-09 | 247 | 420 | 320 | 497 | Trehalase. Trehalase (EC:3.2.1.28) is known to recycle trehalose to glucose. Trehalose is a physiological hallmark of heat-shock response in yeast and protects of proteins and membranes against a variety of stresses. This family is found in conjunction with pfam07492 in fungi. |
| pfam17389 | Bac_rhamnosid6H | 3.93e-07 | 131 | 286 | 77 | 223 | Bacterial alpha-L-rhamnosidase 6 hairpin glycosidase domain. This family consists of bacterial rhamnosidase A and B enzymes. L-Rhamnose is abundant in biomass as a common constituent of glycolipids and glycosides, such as plant pigments, pectic polysaccharides, gums or biosurfactants. Some rhamnosides are important bioactive compounds. For example, terpenyl glycosides, the glycosidic precursor of aromatic terpenoids, act as important flavouring substances in grapes. Other rhamnosides act as cytotoxic rhamnosylated terpenoids, as signal substances in plants or play a role in the antigenicity of pathogenic bacteria. |
| pfam03200 | Glyco_hydro_63 | 7.13e-07 | 168 | 427 | 188 | 492 | Glycosyl hydrolase family 63 C-terminal domain. This is a family of eukaryotic enzymes belonging to glycosyl hydrolase family 63. They catalyze the specific cleavage of the non-reducing terminal glucose residue from Glc(3)Man(9)GlcNAc(2). Mannosyl oligosaccharide glucosidase EC:3.2.1.106 is the first enzyme in the N-linked oligosaccharide processing pathway. This family represents the C-terminal catalytic domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AIQ57834.1 | 5.13e-179 | 4 | 493 | 3 | 484 |
| QHT63246.1 | 6.16e-174 | 4 | 486 | 26 | 503 |
| QIN83484.1 | 1.20e-172 | 3 | 495 | 6 | 498 |
| QHT63247.1 | 1.18e-168 | 4 | 504 | 24 | 521 |
| BBL10718.1 | 3.17e-156 | 4 | 488 | 34 | 509 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P94250 | 1.02e-30 | 92 | 473 | 59 | 431 | Uncharacterized protein BB_0381 OS=Borreliella burgdorferi (strain ATCC 35210 / DSM 4680 / CIP 102532 / B31) OX=224326 GN=BB_0381 PE=4 SV=2 |
| O14255 | 6.09e-13 | 148 | 431 | 533 | 808 | Probable mannosyl-oligosaccharide glucosidase OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPAC6G10.09 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
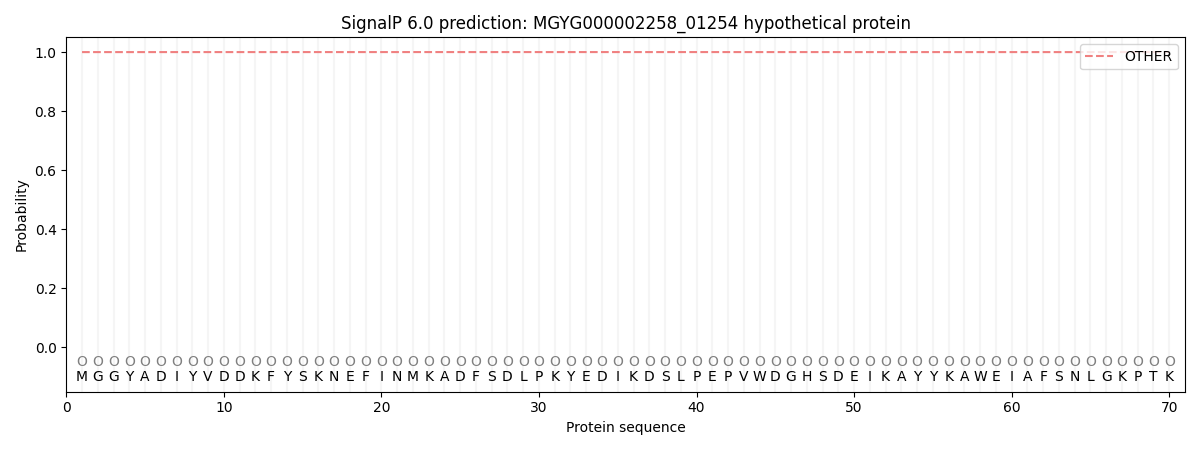
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000052 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
