You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002273_02154
You are here: Home > Sequence: MGYG000002273_02154
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides stercorirosoris | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides stercorirosoris | |||||||||||
| CAZyme ID | MGYG000002273_02154 | |||||||||||
| CAZy Family | GH106 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 21433; End: 24156 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH106 | 32 | 904 | 1.5e-217 | 0.9878640776699029 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam17132 | Glyco_hydro_106 | 1.04e-44 | 159 | 615 | 341 | 775 | alpha-L-rhamnosidase. |
| pfam16355 | DUF4982 | 0.006 | 809 | 835 | 2 | 44 | Domain of unknown function (DUF4982). This family is found in the C-terminal of uncharacterized proteins and beta-galactosidases around 680 residues in length from various Bacteroides species. The function of this protein is unknown. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT88280.1 | 0.0 | 1 | 907 | 1 | 909 |
| ALJ60736.1 | 0.0 | 1 | 907 | 1 | 909 |
| QUT44642.1 | 0.0 | 1 | 906 | 1 | 907 |
| ALJ60737.1 | 2.00e-235 | 25 | 903 | 28 | 907 |
| QUT88279.1 | 1.59e-234 | 25 | 903 | 28 | 907 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6Q2F_A | 8.51e-43 | 167 | 904 | 408 | 1139 | Structureof Rhamnosidase from Novosphingobium sp. PP1Y [Novosphingobium sp. PP1Y] |
| 5MQM_A | 8.78e-40 | 184 | 901 | 386 | 1096 | Glycosidehydrolase BT_0986 [Bacteroides thetaiotaomicron],5MQN_A Glycoside hydrolase BT_0986 [Bacteroides thetaiotaomicron] |
| 5MWK_A | 2.03e-39 | 184 | 901 | 386 | 1096 | Glycosidehydrolase BT_0986 [Bacteroides thetaiotaomicron] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| T2KNA8 | 1.67e-29 | 3 | 726 | 6 | 735 | Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22040 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
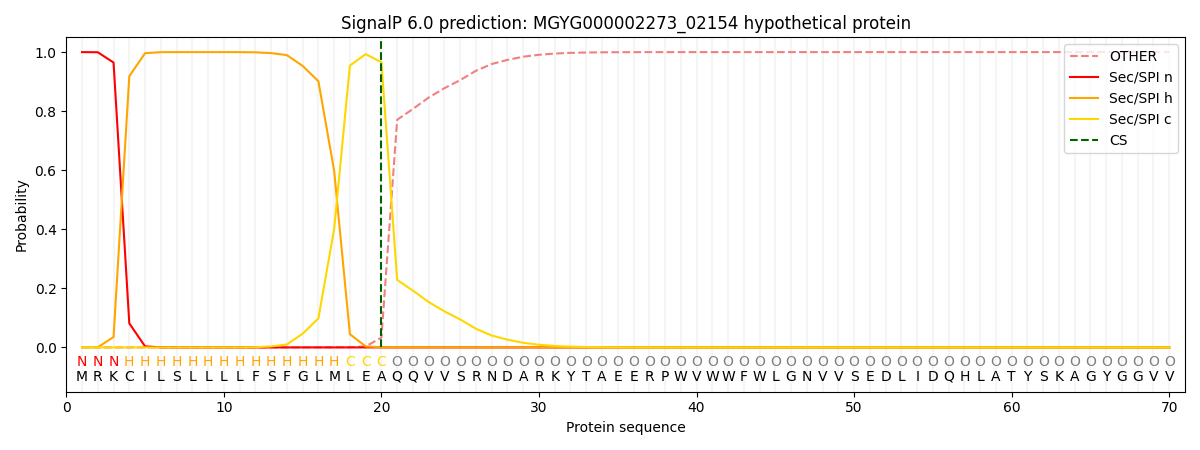
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000331 | 0.999019 | 0.000178 | 0.000156 | 0.000146 | 0.000138 |
