You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002282_00155
You are here: Home > Sequence: MGYG000002282_00155
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
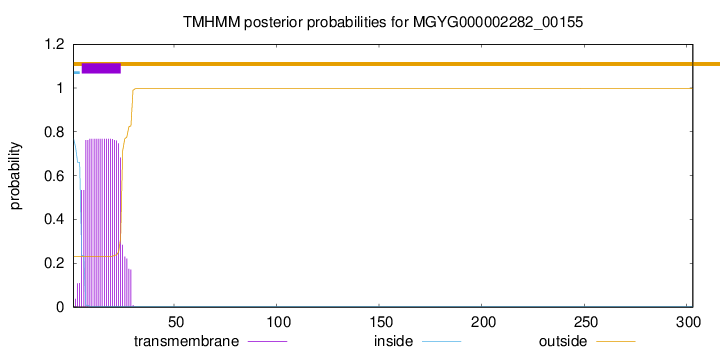
TMHMM annotations
Basic Information help
| Species | Parabacteroides gordonii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Tannerellaceae; Parabacteroides; Parabacteroides gordonii | |||||||||||
| CAZyme ID | MGYG000002282_00155 | |||||||||||
| CAZy Family | GH20 | |||||||||||
| CAZyme Description | Beta-hexosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 203545; End: 204456 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH20 | 135 | 301 | 7.1e-68 | 0.486646884272997 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06563 | GH20_chitobiase-like | 3.61e-100 | 140 | 303 | 1 | 180 | The chitobiase of Serratia marcescens is a beta-N-1,4-acetylhexosaminidase with a glycosyl hydrolase family 20 (GH20) domain that hydrolyzes the beta-1,4-glycosidic linkages in oligomers derived from chitin. Chitin is degraded by a two step process: i) a chitinase hydrolyzes the chitin to oligosaccharides and disaccharides such as di-N-acetyl-D-glucosamine and chitobiose, ii) chitobiase then further degrades these oligomers into monomers. This GH20 domain family includes an N-acetylglucosamidase (GlcNAcase A) from Pseudoalteromonas piscicida and an N-acetylhexosaminidase (SpHex) from Streptomyces plicatus. SpHex lacks the C-terminal PKD (polycystic kidney disease I)-like domain found in the chitobiases. The GH20 hexosaminidases are thought to act via a catalytic mechanism in which the catalytic nucleophile is not provided by solvent or the enzyme, but by the substrate itself. |
| pfam00728 | Glyco_hydro_20 | 2.13e-84 | 140 | 303 | 1 | 175 | Glycosyl hydrolase family 20, catalytic domain. This domain has a TIM barrel fold. |
| cd06568 | GH20_SpHex_like | 1.41e-69 | 140 | 300 | 1 | 166 | A subgroup of the Glycosyl hydrolase family 20 (GH20) catalytic domain found in proteins similar to the N-acetylhexosaminidase from Streptomyces plicatus (SpHex). SpHex catalyzes the hydrolysis of N-acetyl-beta-hexosaminides. An Asp residue within the active site plays a critical role in substrate-assisted catalysis by orienting the 2-acetamido group and stabilizing the transition state. The GH20 hexosaminidases are thought to act via a catalytic mechanism in which the catalytic nucleophile is not provided by solvent or the enzyme, but by the substrate itself. Proteins belonging to this subgroup lack the C-terminal PKD (polycystic kidney disease I)-like domain found in the chitobiases. |
| COG3525 | Chb | 2.48e-68 | 23 | 299 | 136 | 437 | N-acetyl-beta-hexosaminidase [Carbohydrate transport and metabolism]. |
| cd06570 | GH20_chitobiase-like_1 | 1.66e-67 | 140 | 301 | 1 | 160 | A functionally uncharacterized subgroup of the Glycosyl hydrolase family 20 (GH20) catalytic domain found in proteins similar to the chitobiase of Serratia marcescens, a beta-N-1,4-acetylhexosaminidase that hydrolyzes the beta-1,4-glycosidic linkages in oligomers derived from chitin. Chitin is degraded by a two step process: i) a chitinase hydrolyzes the chitin to oligosaccharides and disaccharides such as di-N-acetyl-D-glucosamine and chitobiose, ii) chitobiase then further degrades these oligomers into monomers. This subgroup lacks the C-terminal PKD (polycystic kidney disease I)-like domain found in the chitobiases. The GH20 hexosaminidases are thought to act via a catalytic mechanism in which the catalytic nucleophile is not provided by solvent or the enzyme, but by the substrate itself. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT50216.1 | 1.08e-149 | 5 | 303 | 6 | 304 |
| BAD47130.1 | 9.78e-122 | 21 | 303 | 34 | 291 |
| ANQ62643.1 | 2.78e-121 | 21 | 303 | 34 | 291 |
| CAH06100.1 | 2.78e-121 | 21 | 303 | 34 | 291 |
| QCT76948.1 | 2.78e-121 | 21 | 303 | 34 | 291 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7DUP_A | 1.01e-59 | 25 | 298 | 8 | 316 | ChainA, Beta-N-acetylhexosaminidase [Bacteroides thetaiotaomicron],7DVA_A Chain A, Beta-N-acetylhexosaminidase [Bacteroides thetaiotaomicron],7DVA_B Chain B, Beta-N-acetylhexosaminidase [Bacteroides thetaiotaomicron] |
| 7DVB_A | 5.47e-59 | 25 | 298 | 8 | 316 | ChainA, Beta-N-acetylhexosaminidase [Bacteroides thetaiotaomicron],7DVB_B Chain B, Beta-N-acetylhexosaminidase [Bacteroides thetaiotaomicron],7DVB_C Chain C, Beta-N-acetylhexosaminidase [Bacteroides thetaiotaomicron],7DVB_D Chain D, Beta-N-acetylhexosaminidase [Bacteroides thetaiotaomicron] |
| 6EZR_A | 2.98e-58 | 15 | 302 | 135 | 442 | Crystalstructure of GH20 Exo beta-N-Acetylglucosaminidase from Vibrio harveyi [Vibrio harveyi],6EZR_B Crystal structure of GH20 Exo beta-N-Acetylglucosaminidase from Vibrio harveyi [Vibrio harveyi],6EZS_A Crystal structure of GH20 Exo beta-N-Acetylglucosaminidase from Vibrio harveyi in complex with N-acetylglucosamine [Vibrio harveyi],6EZS_B Crystal structure of GH20 Exo beta-N-Acetylglucosaminidase from Vibrio harveyi in complex with N-acetylglucosamine [Vibrio harveyi],6K35_A Crystal structure of GH20 exo beta-N-acetylglucosaminidase from Vibrio harveyi in complex with NAG-thiazoline [Vibrio harveyi],6K35_B Crystal structure of GH20 exo beta-N-acetylglucosaminidase from Vibrio harveyi in complex with NAG-thiazoline [Vibrio harveyi] |
| 6EZT_A | 4.03e-57 | 15 | 302 | 132 | 439 | Crystalstructure of GH20 Exo beta-N-Acetylglucosaminidase D437A inactive mutant from Vibrio harveyi [Vibrio harveyi],6EZT_B Crystal structure of GH20 Exo beta-N-Acetylglucosaminidase D437A inactive mutant from Vibrio harveyi [Vibrio harveyi] |
| 3RCN_A | 1.47e-55 | 72 | 299 | 63 | 317 | CrystalStructure of Beta-N-Acetylhexosaminidase from Arthrobacter aurescens [Paenarthrobacter aurescens TC1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P96155 | 6.79e-54 | 19 | 302 | 137 | 439 | Beta-hexosaminidase OS=Vibrio furnissii OX=29494 GN=exoI PE=1 SV=1 |
| B2UQG6 | 9.36e-54 | 25 | 298 | 34 | 327 | Beta-hexosaminidase Amuc_0868 OS=Akkermansia muciniphila (strain ATCC BAA-835 / DSM 22959 / JCM 33894 / BCRC 81048 / CCUG 64013 / CIP 107961 / Muc) OX=349741 GN=Amuc_0868 PE=1 SV=1 |
| P49008 | 9.06e-53 | 25 | 298 | 39 | 335 | Beta-hexosaminidase OS=Porphyromonas gingivalis (strain ATCC BAA-308 / W83) OX=242619 GN=nahA PE=3 SV=2 |
| Q04786 | 2.55e-42 | 25 | 298 | 202 | 519 | Beta-hexosaminidase OS=Vibrio vulnificus OX=672 GN=hex PE=3 SV=1 |
| Q7WUL4 | 7.25e-38 | 75 | 299 | 67 | 299 | Beta-N-acetylhexosaminidase OS=Cellulomonas fimi OX=1708 GN=hex20 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
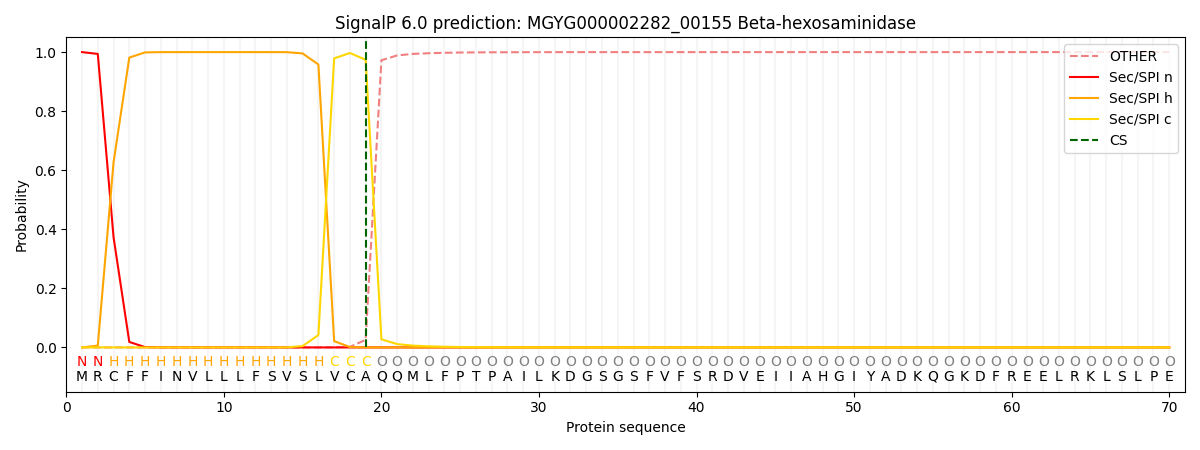
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000283 | 0.999075 | 0.000190 | 0.000138 | 0.000144 | 0.000135 |