You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002308_00908
You are here: Home > Sequence: MGYG000002308_00908
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Campylobacter lanienae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Campylobacterota; Campylobacteria; Campylobacterales; Campylobacteraceae; Campylobacter; Campylobacter lanienae | |||||||||||
| CAZyme ID | MGYG000002308_00908 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | Beta-hexosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 891369; End: 892427 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 86 | 306 | 1.2e-52 | 0.9629629629629629 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1472 | BglX | 5.33e-68 | 23 | 351 | 3 | 314 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| pfam00933 | Glyco_hydro_3 | 2.64e-65 | 23 | 346 | 2 | 314 | Glycosyl hydrolase family 3 N terminal domain. |
| PRK05337 | PRK05337 | 2.97e-51 | 30 | 306 | 3 | 279 | beta-hexosaminidase; Provisional |
| PRK15098 | PRK15098 | 8.07e-09 | 27 | 349 | 51 | 353 | beta-glucosidase BglX. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ARQ97648.1 | 1.18e-250 | 1 | 352 | 1 | 352 |
| ARR00767.1 | 2.04e-219 | 1 | 352 | 1 | 352 |
| ARR04023.1 | 1.09e-207 | 1 | 352 | 1 | 352 |
| ARR02393.1 | 2.56e-206 | 1 | 352 | 1 | 352 |
| ARQ99020.1 | 8.54e-205 | 1 | 352 | 1 | 352 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6K5J_A | 2.37e-50 | 17 | 351 | 7 | 339 | Structureof a glycoside hydrolase family 3 beta-N-acetylglucosaminidase from Paenibacillus sp. str. FPU-7 [Paenibacillaceae] |
| 3BMX_A | 1.06e-41 | 22 | 351 | 43 | 395 | Beta-N-hexosaminidase(YbbD) from Bacillus subtilis [Bacillus subtilis],3BMX_B Beta-N-hexosaminidase (YbbD) from Bacillus subtilis [Bacillus subtilis],3NVD_A Structure of YBBD in complex with pugnac [Bacillus subtilis],3NVD_B Structure of YBBD in complex with pugnac [Bacillus subtilis] |
| 3LK6_A | 4.16e-41 | 22 | 351 | 17 | 369 | ChainA, Lipoprotein ybbD [Bacillus subtilis],3LK6_B Chain B, Lipoprotein ybbD [Bacillus subtilis],3LK6_C Chain C, Lipoprotein ybbD [Bacillus subtilis],3LK6_D Chain D, Lipoprotein ybbD [Bacillus subtilis] |
| 4GYJ_A | 5.49e-41 | 22 | 351 | 47 | 399 | Crystalstructure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1) [Bacillus subtilis subsp. subtilis str. 168],4GYJ_B Crystal structure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1) [Bacillus subtilis subsp. subtilis str. 168],4GYK_A Crystal structure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1211) [Bacillus subtilis subsp. subtilis str. 168],4GYK_B Crystal structure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1211) [Bacillus subtilis subsp. subtilis str. 168] |
| 3TEV_A | 1.11e-35 | 28 | 304 | 19 | 290 | Thecrystal structure of glycosyl hydrolase from Deinococcus radiodurans R1 [Deinococcus radiodurans R1],3TEV_B The crystal structure of glycosyl hydrolase from Deinococcus radiodurans R1 [Deinococcus radiodurans R1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P40406 | 5.83e-41 | 22 | 351 | 43 | 395 | Beta-hexosaminidase OS=Bacillus subtilis (strain 168) OX=224308 GN=nagZ PE=1 SV=1 |
| Q0HVQ8 | 3.34e-38 | 52 | 306 | 21 | 279 | Beta-hexosaminidase OS=Shewanella sp. (strain MR-7) OX=60481 GN=nagZ PE=3 SV=1 |
| Q0HJG7 | 1.27e-37 | 52 | 306 | 21 | 279 | Beta-hexosaminidase OS=Shewanella sp. (strain MR-4) OX=60480 GN=nagZ PE=3 SV=1 |
| Q0AF74 | 2.01e-37 | 49 | 306 | 20 | 282 | Beta-hexosaminidase OS=Nitrosomonas eutropha (strain DSM 101675 / C91 / Nm57) OX=335283 GN=nagZ PE=3 SV=1 |
| Q8EEW2 | 2.47e-37 | 52 | 306 | 21 | 279 | Beta-hexosaminidase OS=Shewanella oneidensis (strain MR-1) OX=211586 GN=nagZ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
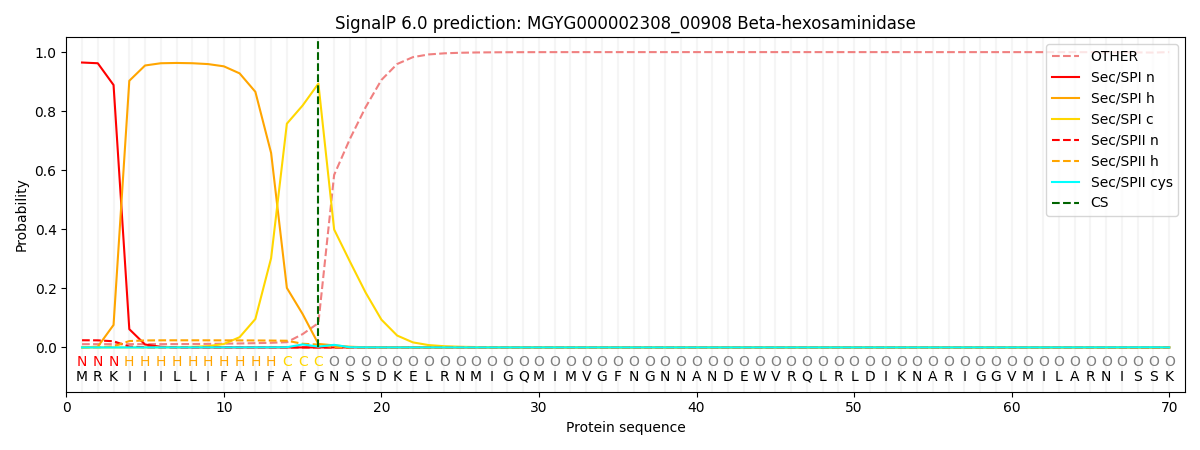
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.016226 | 0.956136 | 0.026638 | 0.000307 | 0.000327 | 0.000352 |
